You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004585_01603
You are here: Home > Sequence: MGYG000004585_01603
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Methanobacterium sp000499765 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Archaea; Methanobacteriota; Methanobacteria; Methanobacteriales; Methanobacteriaceae; Methanobacterium; Methanobacterium sp000499765 | |||||||||||
| CAZyme ID | MGYG000004585_01603 | |||||||||||
| CAZy Family | GT28 | |||||||||||
| CAZyme Description | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 39068; End: 40213 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT28 | 219 | 350 | 2.3e-19 | 0.8662420382165605 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd03785 | GT28_MurG | 8.90e-37 | 2 | 356 | 1 | 337 | undecaprenyldiphospho-muramoylpentapeptide beta-N-acetylglucosaminyltransferase. MurG (EC 2.4.1.227) is an N-acetylglucosaminyltransferase, the last enzyme involved in the intracellular phase of peptidoglycan biosynthesis. It transfers N-acetyl-D-glucosamine (GlcNAc) from UDP-GlcNAc to the C4 hydroxyl of a lipid-linked N-acetylmuramoyl pentapeptide (NAM). The resulting disaccharide is then transported across the cell membrane, where it is polymerized into NAG-NAM cell-wall repeat structure. MurG belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains, each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
| COG1819 | YjiC | 1.25e-29 | 1 | 381 | 2 | 402 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
| TIGR00661 | MJ1255 | 2.21e-28 | 2 | 326 | 1 | 296 | conserved hypothetical protein. This model represents nearly the full length of MJ1255 from Methanococcus jannaschii and of an unpublished protein from Vibrio cholerae, as well as the C-terminal half of a protein from Methanobacterium thermoautotrophicum. A small region (~50 amino acids) within the domain appears related to a family of sugar transferases. [Hypothetical proteins, Conserved] |
| COG0707 | MurG | 9.19e-27 | 1 | 380 | 1 | 357 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
| PRK00726 | murG | 7.51e-21 | 13 | 380 | 13 | 357 | undecaprenyldiphospho-muramoylpentapeptide beta-N- acetylglucosaminyltransferase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CDG65339.1 | 1.50e-279 | 1 | 381 | 1 | 381 |
| AXV40963.1 | 8.41e-261 | 1 | 381 | 1 | 381 |
| AIS31274.1 | 2.31e-258 | 1 | 381 | 1 | 381 |
| CEA12741.1 | 2.31e-258 | 1 | 381 | 1 | 381 |
| CEL25122.1 | 2.31e-258 | 1 | 381 | 1 | 381 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8TWQ5 | 2.56e-37 | 19 | 323 | 1 | 295 | Uncharacterized glycosyltransferase MK0977 OS=Methanopyrus kandleri (strain AV19 / DSM 6324 / JCM 9639 / NBRC 100938) OX=190192 GN=MK0977 PE=3 SV=1 |
| Q8PWF3 | 1.01e-35 | 1 | 375 | 1 | 371 | Uncharacterized glycosyltransferase MM_1636 OS=Methanosarcina mazei (strain ATCC BAA-159 / DSM 3647 / Goe1 / Go1 / JCM 11833 / OCM 88) OX=192952 GN=MM_1636 PE=3 SV=1 |
| Q8TTI1 | 1.36e-32 | 1 | 331 | 1 | 328 | Uncharacterized glycosyltransferase MA_0452 OS=Methanosarcina acetivorans (strain ATCC 35395 / DSM 2834 / JCM 12185 / C2A) OX=188937 GN=MA_0452 PE=3 SV=1 |
| Q58652 | 2.94e-29 | 1 | 374 | 1 | 362 | Uncharacterized glycosyltransferase MJ1255 OS=Methanocaldococcus jannaschii (strain ATCC 43067 / DSM 2661 / JAL-1 / JCM 10045 / NBRC 100440) OX=243232 GN=MJ1255 PE=3 SV=2 |
| Q8PZB2 | 2.00e-27 | 1 | 325 | 1 | 322 | Uncharacterized glycosyltransferase MM_0582 OS=Methanosarcina mazei (strain ATCC BAA-159 / DSM 3647 / Goe1 / Go1 / JCM 11833 / OCM 88) OX=192952 GN=MM_0582 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
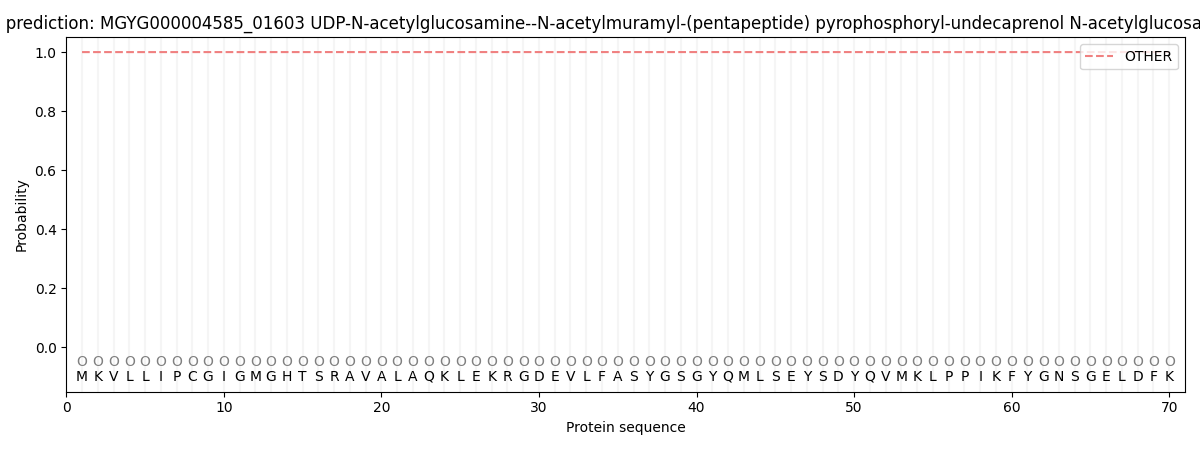
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000062 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
