You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004290_02637
You are here: Home > Sequence: MGYG000004290_02637
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes sp900549305 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes sp900549305 | |||||||||||
| CAZyme ID | MGYG000004290_02637 | |||||||||||
| CAZy Family | CE9 | |||||||||||
| CAZyme Description | N-acetylglucosamine-6-phosphate deacetylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 57; End: 1307 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE9 | 28 | 410 | 1.4e-107 | 0.9946380697050938 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00854 | NagA | 9.15e-133 | 26 | 410 | 1 | 374 | N-acetylglucosamine-6-phosphate deacetylase, NagA, catalyzes the hydrolysis of the N-acetyl group of N-acetyl-glucosamine-6-phosphate (GlcNAc-6-P) to glucosamine 6-phosphate and acetate. This is the first committed step in the biosynthetic pathway to amino-sugar-nucleotides, which is needed for cell wall peptidoglycan and teichoic acid biosynthesis. Deacetylation of N-acetylglucosamine is also important in lipopolysaccharide synthesis and cell wall recycling. |
| COG1820 | NagA | 5.74e-110 | 27 | 415 | 4 | 380 | N-acetylglucosamine-6-phosphate deacetylase [Carbohydrate transport and metabolism]. |
| TIGR00221 | nagA | 5.59e-78 | 22 | 411 | 2 | 380 | N-acetylglucosamine-6-phosphate deacetylase. [Central intermediary metabolism, Amino sugars] |
| PRK11170 | nagA | 5.48e-51 | 38 | 413 | 15 | 379 | N-acetylglucosamine-6-phosphate deacetylase; Provisional |
| cd01297 | D-aminoacylase | 1.32e-23 | 42 | 415 | 20 | 404 | D-aminoacylases (N-acyl-D-Amino acid amidohydrolases) catalyze the hydrolysis of N-acyl-D-amino acids to produce the corresponding D-amino acids, which are used as intermediates in the synthesis of pesticides, bioactive peptides, and antibiotics. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI64809.1 | 4.27e-154 | 24 | 416 | 3 | 393 |
| QRX64347.1 | 2.64e-150 | 23 | 416 | 3 | 396 |
| SCD19170.1 | 8.67e-149 | 23 | 416 | 3 | 396 |
| AYQ35214.1 | 5.93e-148 | 24 | 416 | 2 | 391 |
| AEI48316.1 | 2.40e-147 | 24 | 416 | 2 | 391 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6FV3_A | 1.48e-45 | 77 | 411 | 66 | 390 | Crystalstructure of N-acetyl-D-glucosamine-6-phosphate deacetylase from Mycobacterium smegmatis. [Mycolicibacterium smegmatis MC2 155],6FV3_B Crystal structure of N-acetyl-D-glucosamine-6-phosphate deacetylase from Mycobacterium smegmatis. [Mycolicibacterium smegmatis MC2 155],6FV3_C Crystal structure of N-acetyl-D-glucosamine-6-phosphate deacetylase from Mycobacterium smegmatis. [Mycolicibacterium smegmatis MC2 155],6FV3_D Crystal structure of N-acetyl-D-glucosamine-6-phosphate deacetylase from Mycobacterium smegmatis. [Mycolicibacterium smegmatis MC2 155] |
| 1O12_A | 1.67e-45 | 77 | 416 | 55 | 375 | Crystalstructure of N-acetylglucosamine-6-phosphate deacetylase (TM0814) from Thermotoga maritima at 2.5 A resolution [Thermotoga maritima],1O12_B Crystal structure of N-acetylglucosamine-6-phosphate deacetylase (TM0814) from Thermotoga maritima at 2.5 A resolution [Thermotoga maritima] |
| 6FV4_A | 2.11e-44 | 77 | 411 | 66 | 390 | Thestructure of N-acetyl-D-glucosamine-6-phosphate deacetylase D267A mutant from Mycobacterium smegmatis in complex with N-acetyl-D-glucosamine-6-phosphate [Mycolicibacterium smegmatis MC2 155],6FV4_B The structure of N-acetyl-D-glucosamine-6-phosphate deacetylase D267A mutant from Mycobacterium smegmatis in complex with N-acetyl-D-glucosamine-6-phosphate [Mycolicibacterium smegmatis MC2 155] |
| 2VHL_A | 1.42e-42 | 31 | 414 | 11 | 388 | TheThree-dimensional structure of the N-Acetylglucosamine-6- phosphate deacetylase from Bacillus subtilis [Bacillus subtilis],2VHL_B The Three-dimensional structure of the N-Acetylglucosamine-6- phosphate deacetylase from Bacillus subtilis [Bacillus subtilis] |
| 7NUT_A | 1.32e-38 | 69 | 415 | 57 | 403 | ChainA, N-acetylglucosamine-6-phosphate deacetylase [Homo sapiens],7NUT_B Chain B, N-acetylglucosamine-6-phosphate deacetylase [Homo sapiens],7NUU_A Chain A, N-acetylglucosamine-6-phosphate deacetylase [Homo sapiens],7NUU_B Chain B, N-acetylglucosamine-6-phosphate deacetylase [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q84F86 | 5.59e-47 | 29 | 412 | 10 | 381 | N-acetylglucosamine-6-phosphate deacetylase OS=Lysinibacillus sphaericus OX=1421 GN=nagA PE=2 SV=1 |
| Q6P0U0 | 6.47e-43 | 68 | 416 | 56 | 404 | N-acetylglucosamine-6-phosphate deacetylase OS=Danio rerio OX=7955 GN=amdhd2 PE=2 SV=1 |
| O34450 | 7.76e-42 | 31 | 414 | 11 | 388 | N-acetylglucosamine-6-phosphate deacetylase OS=Bacillus subtilis (strain 168) OX=224308 GN=nagA PE=1 SV=1 |
| Q8XAC3 | 1.01e-41 | 46 | 411 | 24 | 373 | N-acetylgalactosamine-6-phosphate deacetylase OS=Escherichia coli O157:H7 OX=83334 GN=agaA PE=1 SV=2 |
| P44537 | 5.83e-40 | 45 | 411 | 22 | 377 | N-acetylglucosamine-6-phosphate deacetylase OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=nagA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
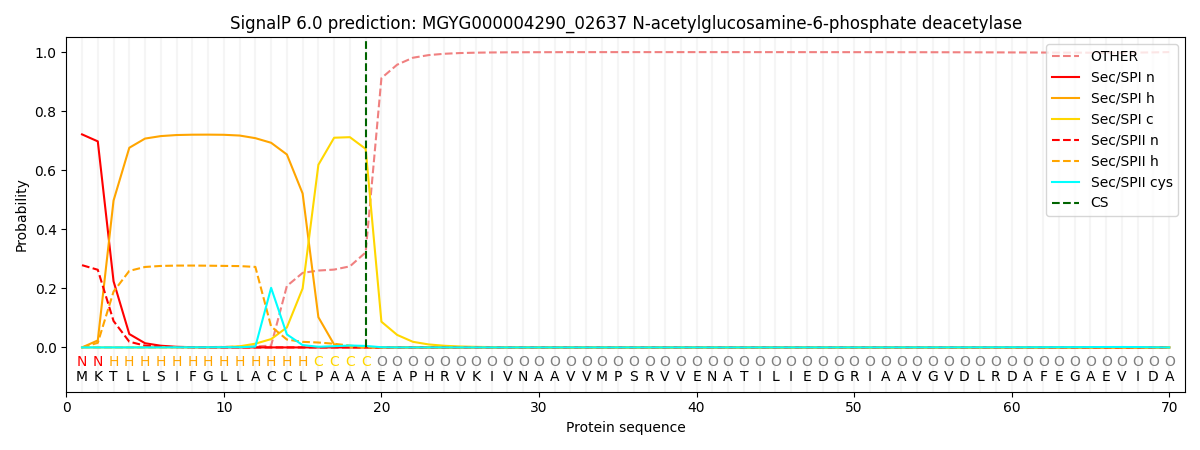
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000590 | 0.710811 | 0.287708 | 0.000340 | 0.000285 | 0.000241 |
