You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003941_00165
You are here: Home > Sequence: MGYG000003941_00165
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; | |||||||||||
| CAZyme ID | MGYG000003941_00165 | |||||||||||
| CAZy Family | GH13 | |||||||||||
| CAZyme Description | Alpha-amylase 2 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2634; End: 4025 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH13 | 50 | 329 | 8.5e-57 | 0.9297658862876255 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd11313 | AmyAc_arch_bac_AmyA | 4.30e-170 | 33 | 368 | 1 | 336 | Alpha amylase catalytic domain found in archaeal and bacterial Alpha-amylases (also called 1,4-alpha-D-glucan-4-glucanohydrolase). AmyA (EC 3.2.1.1) catalyzes the hydrolysis of alpha-(1,4) glycosidic linkages of glycogen, starch, related polysaccharides, and some oligosaccharides. This group includes firmicutes, bacteroidetes, and proteobacteria. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| cd11316 | AmyAc_bac2_AmyA | 3.19e-52 | 60 | 368 | 29 | 403 | Alpha amylase catalytic domain found in bacterial Alpha-amylases (also called 1,4-alpha-D-glucan-4-glucanohydrolase). AmyA (EC 3.2.1.1) catalyzes the hydrolysis of alpha-(1,4) glycosidic linkages of glycogen, starch, related polysaccharides, and some oligosaccharides. This group includes Chloroflexi, Dictyoglomi, and Fusobacteria. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| cd11338 | AmyAc_CMD | 3.17e-46 | 51 | 369 | 53 | 388 | Alpha amylase catalytic domain found in cyclomaltodextrinases and related proteins. Cyclomaltodextrinase (CDase; EC3.2.1.54), neopullulanase (NPase; EC 3.2.1.135), and maltogenic amylase (MA; EC 3.2.1.133) catalyze the hydrolysis of alpha-(1,4) glycosidic linkages on a number of substrates including cyclomaltodextrins (CDs), pullulan, and starch. These enzymes hydrolyze CDs and starch to maltose and pullulan to panose by cleavage of alpha-1,4 glycosidic bonds whereas alpha-amylases essentially lack activity on CDs and pullulan. They also catalyze transglycosylation of oligosaccharides to the C3-, C4- or C6-hydroxyl groups of various acceptor sugar molecules. Since these proteins are nearly indistinguishable from each other, they are referred to as cyclomaltodextrinases (CMDs). The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| cd11333 | AmyAc_SI_OligoGlu_DGase | 1.12e-44 | 60 | 361 | 31 | 428 | Alpha amylase catalytic domain found in Sucrose isomerases, oligo-1,6-glucosidase (also called isomaltase; sucrase-isomaltase; alpha-limit dextrinase), dextran glucosidase (also called glucan 1,6-alpha-glucosidase), and related proteins. The sucrose isomerases (SIs) Isomaltulose synthase (EC 5.4.99.11) and Trehalose synthase (EC 5.4.99.16) catalyze the isomerization of sucrose and maltose to produce isomaltulose and trehalulose, respectively. Oligo-1,6-glucosidase (EC 3.2.1.10) hydrolyzes the alpha-1,6-glucosidic linkage of isomaltooligosaccharides, pannose, and dextran. Unlike alpha-1,4-glucosidases (EC 3.2.1.20), it fails to hydrolyze the alpha-1,4-glucosidic bonds of maltosaccharides. Dextran glucosidase (DGase, EC 3.2.1.70) hydrolyzes alpha-1,6-glucosidic linkages at the non-reducing end of panose, isomaltooligosaccharides and dextran to produce alpha-glucose.The common reaction chemistry of the alpha-amylase family enzymes is based on a two-step acid catalytic mechanism that requires two critical carboxylates: one acting as a general acid/base (Glu) and the other as a nucleophile (Asp). Both hydrolysis and transglycosylation proceed via the nucleophilic substitution reaction between the anomeric carbon, C1 and a nucleophile. Both enzymes contain the three catalytic residues (Asp, Glu and Asp) common to the alpha-amylase family as well as two histidine residues which are predicted to be critical to binding the glucose residue adjacent to the scissile bond in the substrates. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| COG0366 | AmyA | 1.01e-43 | 41 | 409 | 16 | 488 | Glycosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFL78249.1 | 1.02e-238 | 1 | 444 | 1 | 440 |
| CBK62785.1 | 2.86e-238 | 14 | 444 | 3 | 430 |
| BBL08213.1 | 5.87e-238 | 1 | 444 | 1 | 440 |
| BBL11004.1 | 5.87e-238 | 1 | 444 | 1 | 440 |
| BBL00132.1 | 1.57e-204 | 1 | 437 | 1 | 416 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3DHU_A | 6.21e-57 | 38 | 359 | 14 | 347 | Crystalstructure of an alpha-amylase from Lactobacillus plantarum [Lactiplantibacillus plantarum],3DHU_B Crystal structure of an alpha-amylase from Lactobacillus plantarum [Lactiplantibacillus plantarum],3DHU_C Crystal structure of an alpha-amylase from Lactobacillus plantarum [Lactiplantibacillus plantarum],3DHU_D Crystal structure of an alpha-amylase from Lactobacillus plantarum [Lactiplantibacillus plantarum] |
| 4GKL_A | 5.47e-53 | 38 | 412 | 7 | 376 | Crystalstructure of a noncanonic maltogenic alpha-amylase AmyB from Thermotoga neapolitana [Thermotoga neapolitana],4GKL_B Crystal structure of a noncanonic maltogenic alpha-amylase AmyB from Thermotoga neapolitana [Thermotoga neapolitana] |
| 2ZE0_A | 1.97e-32 | 33 | 410 | 5 | 512 | Alpha-glucosidaseGSJ [Geobacillus sp. HTA-462] |
| 5H2T_A | 2.14e-29 | 31 | 405 | 20 | 511 | Structureof trehalose synthase [Thermomonospora curvata DSM 43183],5H2T_B Structure of trehalose synthase [Thermomonospora curvata DSM 43183],5H2T_C Structure of trehalose synthase [Thermomonospora curvata DSM 43183],5H2T_D Structure of trehalose synthase [Thermomonospora curvata DSM 43183],5H2T_E Structure of trehalose synthase [Thermomonospora curvata DSM 43183],5H2T_F Structure of trehalose synthase [Thermomonospora curvata DSM 43183],5H2T_G Structure of trehalose synthase [Thermomonospora curvata DSM 43183],5H2T_H Structure of trehalose synthase [Thermomonospora curvata DSM 43183] |
| 3WY3_A | 6.20e-27 | 33 | 439 | 7 | 530 | Crystalstructure of alpha-glucosidase mutant D202N in complex with glucose and glycerol [Halomonas sp. H11],3WY3_B Crystal structure of alpha-glucosidase mutant D202N in complex with glucose and glycerol [Halomonas sp. H11] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| L8B068 | 1.43e-34 | 31 | 406 | 242 | 614 | Alpha-amylase MalA OS=Haloarcula japonica (strain ATCC 49778 / DSM 6131 / JCM 7785 / NBRC 101032 / NCIMB 13157 / TR-1) OX=1227453 GN=malA PE=1 SV=1 |
| P20845 | 1.73e-28 | 6 | 407 | 9 | 482 | Alpha-amylase OS=Priestia megaterium OX=1404 PE=1 SV=1 |
| O06458 | 3.93e-28 | 31 | 444 | 3 | 535 | Trehalose synthase OS=Thermus thermophilus OX=274 GN=treS PE=3 SV=1 |
| P14898 | 4.28e-28 | 31 | 425 | 128 | 540 | Alpha-amylase 2 OS=Dictyoglomus thermophilum (strain ATCC 35947 / DSM 3960 / H-6-12) OX=309799 GN=amyB PE=1 SV=2 |
| P14899 | 1.23e-27 | 33 | 329 | 31 | 355 | Alpha-amylase 3 OS=Dictyoglomus thermophilum (strain ATCC 35947 / DSM 3960 / H-6-12) OX=309799 GN=amyC PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
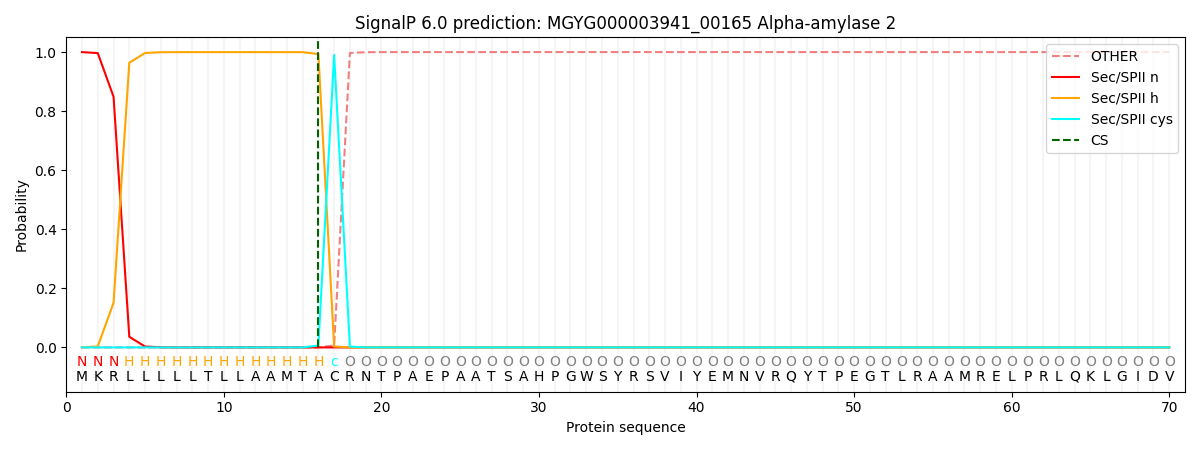
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000047 | 0.999989 | 0.000000 | 0.000000 | 0.000000 |
