You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003710_01030
You are here: Home > Sequence: MGYG000003710_01030
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Loigolactobacillus coryniformis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Loigolactobacillus; Loigolactobacillus coryniformis | |||||||||||
| CAZyme ID | MGYG000003710_01030 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | N-acetylglucosaminyl-diphospho-decaprenol L-rhamnosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1816; End: 2595 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd04186 | GT_2_like_c | 2.13e-34 | 6 | 229 | 1 | 166 | Subfamily of Glycosyltransferase Family GT2 of unknown function. GT-2 includes diverse families of glycosyltransferases with a common GT-A type structural fold, which has two tightly associated beta/alpha/beta domains that tend to form a continuous central sheet of at least eight beta-strands. These are enzymes that catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. Glycosyltransferases have been classified into more than 90 distinct sequence based families. |
| COG1216 | GT2 | 3.78e-29 | 1 | 253 | 2 | 245 | Glycosyltransferase, GT2 family [Carbohydrate transport and metabolism]. |
| cd02525 | Succinoglycan_BP_ExoA | 1.34e-07 | 176 | 250 | 157 | 223 | ExoA is involved in the biosynthesis of succinoglycan. Succinoglycan Biosynthesis Protein ExoA catalyzes the formation of a beta-1,3 linkage of the second sugar (glucose) of the succinoglycan with the galactose on the lipid carrie. Succinoglycan is an acidic exopolysaccharide that is important for invasion of the nodules. Succinoglycan is a high-molecular-weight polymer composed of repeating octasaccharide units. These units are synthesized on membrane-bound isoprenoid lipid carriers, beginning with galactose followed by seven glucose molecules, and modified by the addition of acetate, succinate, and pyruvate. ExoA is a membrane protein with a transmembrance domain at c-terminus. |
| pfam13641 | Glyco_tranf_2_3 | 1.64e-07 | 176 | 238 | 161 | 222 | Glycosyltransferase like family 2. Members of this family of prokaryotic proteins include putative glucosyltransferase, which are involved in bacterial capsule biosynthesis. |
| cd02526 | GT2_RfbF_like | 5.76e-07 | 8 | 251 | 3 | 227 | RfbF is a putative dTDP-rhamnosyl transferase. Shigella flexneri RfbF protein is a putative dTDP-rhamnosyl transferase. dTDP rhamnosyl transferases of Shigella flexneri add rhamnose sugars to N-acetyl-glucosamine in the O-antigen tetrasaccharide repeat. Lipopolysaccharide O antigens are important virulence determinants for many bacteria. The variations of sugar composition, the sequence of the sugars and the linkages in the O antigen provide structural diversity of the O antigen. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATO44694.1 | 3.18e-190 | 1 | 259 | 1 | 259 |
| QEA52878.1 | 5.27e-189 | 1 | 259 | 1 | 259 |
| ATO56466.1 | 7.49e-189 | 1 | 259 | 1 | 259 |
| SFZ87249.1 | 1.38e-136 | 1 | 259 | 1 | 259 |
| ANK70903.1 | 7.08e-111 | 4 | 259 | 5 | 259 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P9WMY2 | 3.37e-15 | 32 | 233 | 32 | 241 | N-acetylglucosaminyl-diphospho-decaprenol L-rhamnosyltransferase OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=wbbL PE=3 SV=2 |
| P9WMY3 | 3.37e-15 | 32 | 233 | 32 | 241 | N-acetylglucosaminyl-diphospho-decaprenol L-rhamnosyltransferase OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=wbbL PE=1 SV=2 |
| P36667 | 3.63e-11 | 69 | 250 | 60 | 243 | Rhamnosyltransferase WbbL OS=Escherichia coli (strain K12) OX=83333 GN=wbbL PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
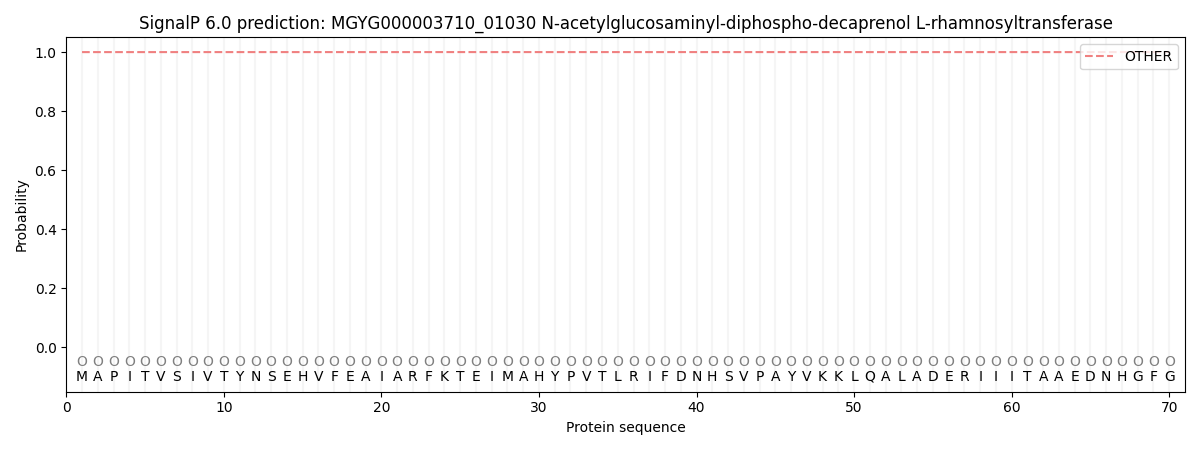
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000046 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
