You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003626_00124
You are here: Home > Sequence: MGYG000003626_00124
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | RC9 sp900771005 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA932; RC9; RC9 sp900771005 | |||||||||||
| CAZyme ID | MGYG000003626_00124 | |||||||||||
| CAZy Family | GH53 | |||||||||||
| CAZyme Description | Arabinogalactan endo-beta-1,4-galactanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3478; End: 4329 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH53 | 28 | 264 | 1.4e-77 | 0.6432748538011696 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07745 | Glyco_hydro_53 | 6.28e-78 | 28 | 261 | 3 | 211 | Glycosyl hydrolase family 53. This domain belongs to family 53 of the glycosyl hydrolase classification. These enzymes are enzymes are endo-1,4- beta-galactanases (EC:3.2.1.89). The structure of this domain is known and has a TIM barrel fold. |
| COG3867 | GanB | 6.48e-67 | 23 | 260 | 37 | 256 | Arabinogalactan endo-1,4-beta-galactosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QJR97948.1 | 1.98e-125 | 25 | 261 | 13 | 249 |
| ADE81415.1 | 4.32e-93 | 4 | 269 | 2 | 243 |
| QVJ79596.1 | 6.94e-89 | 4 | 269 | 2 | 243 |
| SCM59778.1 | 6.10e-87 | 28 | 262 | 42 | 251 |
| ADV45000.1 | 4.10e-85 | 28 | 267 | 49 | 291 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6GPA_A | 1.26e-68 | 22 | 280 | 4 | 239 | Beta-1,4-galactanasefrom Bacteroides thetaiotaomicron with galactose [Bacteroides thetaiotaomicron VPI-5482],6GPA_B Beta-1,4-galactanase from Bacteroides thetaiotaomicron with galactose [Bacteroides thetaiotaomicron VPI-5482] |
| 6GP5_A | 3.78e-68 | 22 | 280 | 40 | 275 | Beta-1,4-galactanasefrom Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],6GP5_B Beta-1,4-galactanase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
| 7OSK_A | 1.46e-36 | 33 | 260 | 61 | 266 | ChainA, Arabinogalactan endo-1,4-beta-galactosidase [Ignisphaera aggregans DSM 17230],7OSK_B Chain B, Arabinogalactan endo-1,4-beta-galactosidase [Ignisphaera aggregans DSM 17230] |
| 1R8L_A | 1.76e-28 | 19 | 260 | 18 | 237 | Thestructure of endo-beta-1,4-galactanase from Bacillus licheniformis [Bacillus licheniformis],1R8L_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis [Bacillus licheniformis],1UR0_A The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR0_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR4_A The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR4_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],2CCR_A Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2CCR_B Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2J74_A Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2J74_B Structure of Beta-1,4-Galactanase [Bacillus licheniformis] |
| 2GFT_A | 1.76e-28 | 19 | 260 | 18 | 237 | ChainA, Glycosyl Hydrolase Family 53 [Bacillus licheniformis],2GFT_B Chain B, Glycosyl Hydrolase Family 53 [Bacillus licheniformis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48843 | 8.24e-46 | 22 | 260 | 3 | 223 | Uncharacterized protein in bgaB 5'region (Fragment) OS=Niallia circulans OX=1397 PE=3 SV=1 |
| Q0CTQ7 | 1.73e-28 | 28 | 264 | 20 | 237 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=galA PE=3 SV=1 |
| A2RB93 | 1.27e-27 | 28 | 264 | 22 | 238 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=galA PE=3 SV=1 |
| Q65CX5 | 1.30e-27 | 19 | 260 | 43 | 262 | Endo-beta-1,4-galactanase OS=Bacillus licheniformis (strain ATCC 14580 / DSM 13 / JCM 2505 / CCUG 7422 / NBRC 12200 / NCIMB 9375 / NCTC 10341 / NRRL NRS-1264 / Gibson 46) OX=279010 GN=ganB PE=1 SV=1 |
| A1D3T4 | 1.39e-27 | 28 | 264 | 27 | 244 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=galA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
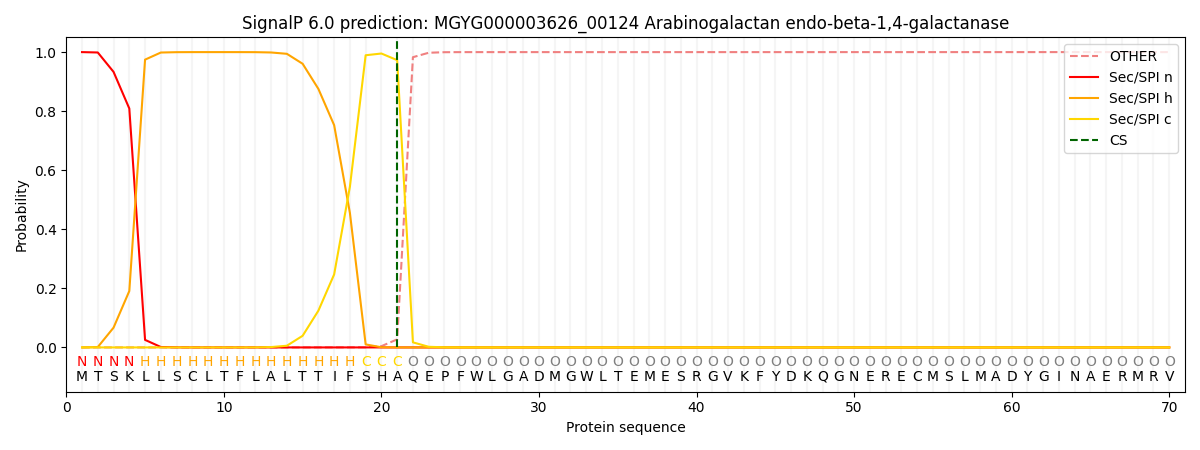
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000277 | 0.998972 | 0.000240 | 0.000168 | 0.000164 | 0.000149 |
