You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003417_01202
You are here: Home > Sequence: MGYG000003417_01202
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Firm-11 sp004556545 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; CAG-74; Firm-11; Firm-11 sp004556545 | |||||||||||
| CAZyme ID | MGYG000003417_01202 | |||||||||||
| CAZy Family | GH29 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 19085; End: 20407 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH29 | 3 | 351 | 3.8e-107 | 0.9219653179190751 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00812 | Alpha_L_fucos | 8.66e-130 | 3 | 380 | 6 | 370 | Alpha-L-fucosidase. O-Glycosyl hydrolases (EC 3.2.1.-) are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycosyl hydrolases, based on sequence similarity, has led to the definition of 85 different families. This classification is available on the CAZy (CArbohydrate-Active EnZymes) web site. Because the fold of proteins is better conserved than their sequences, some of the families can be grouped in 'clans'. Family 29 encompasses alpha-L-fucosidases, which is a lysosomal enzyme responsible for hydrolyzing the alpha-1,6-linked fucose joined to the reducing-end N-acetylglucosamine of the carbohydrate moieties of glycoproteins. Deficiency of alpha-L-fucosidase results in the lysosomal storage disease fucosidosis. |
| pfam01120 | Alpha_L_fucos | 1.13e-126 | 2 | 347 | 4 | 333 | Alpha-L-fucosidase. |
| COG3669 | AfuC | 5.74e-77 | 30 | 381 | 1 | 352 | Alpha-L-fucosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACL33092.1 | 8.48e-182 | 2 | 420 | 7 | 427 |
| QHB43606.1 | 8.48e-182 | 2 | 420 | 7 | 427 |
| AGO17219.1 | 2.42e-181 | 2 | 420 | 7 | 427 |
| AWY44563.1 | 2.42e-181 | 2 | 420 | 7 | 427 |
| ATW44517.1 | 3.43e-181 | 2 | 420 | 7 | 427 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1ODU_A | 4.97e-156 | 3 | 379 | 7 | 386 | CrystalStructure Of Thermotoga Maritima Alpha-Fucosidase In Complex With Fucose [Thermotoga maritima MSB8],1ODU_B Crystal Structure Of Thermotoga Maritima Alpha-Fucosidase In Complex With Fucose [Thermotoga maritima MSB8] |
| 2ZWY_A | 6.09e-156 | 3 | 379 | 7 | 386 | alpha-L-fucosidase[Thermotoga maritima],2ZWY_B alpha-L-fucosidase [Thermotoga maritima],2ZWZ_A alpha-L-fucosidase complexed with inhibitor, Core1 [Thermotoga maritima],2ZWZ_B alpha-L-fucosidase complexed with inhibitor, Core1 [Thermotoga maritima],2ZX5_A alpha-L-fucosidase complexed with inhibitor, F10 [Thermotoga maritima],2ZX5_B alpha-L-fucosidase complexed with inhibitor, F10 [Thermotoga maritima],2ZX6_A alpha-L-fucosidase complexed with inhibitor, F10-1C [Thermotoga maritima],2ZX6_B alpha-L-fucosidase complexed with inhibitor, F10-1C [Thermotoga maritima],2ZX7_A alpha-L-fucosidase complexed with inhibitor, F10-2C [Thermotoga maritima],2ZX7_B alpha-L-fucosidase complexed with inhibitor, F10-2C [Thermotoga maritima],2ZX8_A alpha-L-fucosidase complexed with inhibitor, F10-2C-O [Thermotoga maritima],2ZX8_B alpha-L-fucosidase complexed with inhibitor, F10-2C-O [Thermotoga maritima],2ZX9_A alpha-L-fucosidase complexed with inhibitor, B4 [Thermotoga maritima],2ZX9_B alpha-L-fucosidase complexed with inhibitor, B4 [Thermotoga maritima],2ZXA_A alpha-L-fucosidase complexed with inhibitor, FNJ-acetyl [Thermotoga maritima],2ZXA_B alpha-L-fucosidase complexed with inhibitor, FNJ-acetyl [Thermotoga maritima],2ZXB_A alpha-L-fucosidase complexed with inhibitor, ph-6FNJ [Thermotoga maritima],2ZXB_B alpha-L-fucosidase complexed with inhibitor, ph-6FNJ [Thermotoga maritima],2ZXD_A alpha-L-fucosidase complexed with inhibitor, iso-6FNJ [Thermotoga maritima],2ZXD_B alpha-L-fucosidase complexed with inhibitor, iso-6FNJ [Thermotoga maritima] |
| 1HL8_A | 9.99e-156 | 3 | 379 | 7 | 386 | CrystalStructure Of Thermotoga Maritima Alpha-Fucosidase [Thermotoga maritima MSB8],1HL8_B Crystal Structure Of Thermotoga Maritima Alpha-Fucosidase [Thermotoga maritima MSB8],1HL9_A Crystal Structure Of Thermotoga Maritima Alpha-Fucosidase In Complex With A Mechanism Based Inhibitor [Thermotoga maritima MSB8],1HL9_B Crystal Structure Of Thermotoga Maritima Alpha-Fucosidase In Complex With A Mechanism Based Inhibitor [Thermotoga maritima MSB8] |
| 2WSP_A | 5.71e-155 | 3 | 379 | 7 | 386 | Thermotogamaritima alpha-L-fucosynthase, TmD224G, in complex with alpha-L-Fuc-(1-2)-beta-L-Fuc-N3 [Thermotoga maritima MSB8],2WSP_B Thermotoga maritima alpha-L-fucosynthase, TmD224G, in complex with alpha-L-Fuc-(1-2)-beta-L-Fuc-N3 [Thermotoga maritima MSB8] |
| 7LJJ_A | 6.60e-52 | 12 | 352 | 52 | 402 | ChainA, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135],7LK7_A Chain A, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| C3YWU0 | 6.97e-72 | 3 | 390 | 17 | 393 | Alpha-L-fucosidase OS=Branchiostoma floridae OX=7739 GN=BRAFLDRAFT_56888 PE=3 SV=2 |
| P10901 | 8.57e-69 | 2 | 391 | 19 | 402 | Alpha-L-fucosidase OS=Dictyostelium discoideum OX=44689 GN=alfA PE=3 SV=1 |
| Q2KIM0 | 1.54e-67 | 3 | 380 | 39 | 400 | Tissue alpha-L-fucosidase OS=Bos taurus OX=9913 GN=FUCA1 PE=2 SV=1 |
| Q9BTY2 | 4.16e-67 | 3 | 361 | 33 | 379 | Plasma alpha-L-fucosidase OS=Homo sapiens OX=9606 GN=FUCA2 PE=1 SV=2 |
| Q5RFI5 | 1.10e-66 | 3 | 361 | 31 | 377 | Plasma alpha-L-fucosidase OS=Pongo abelii OX=9601 GN=FUCA2 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
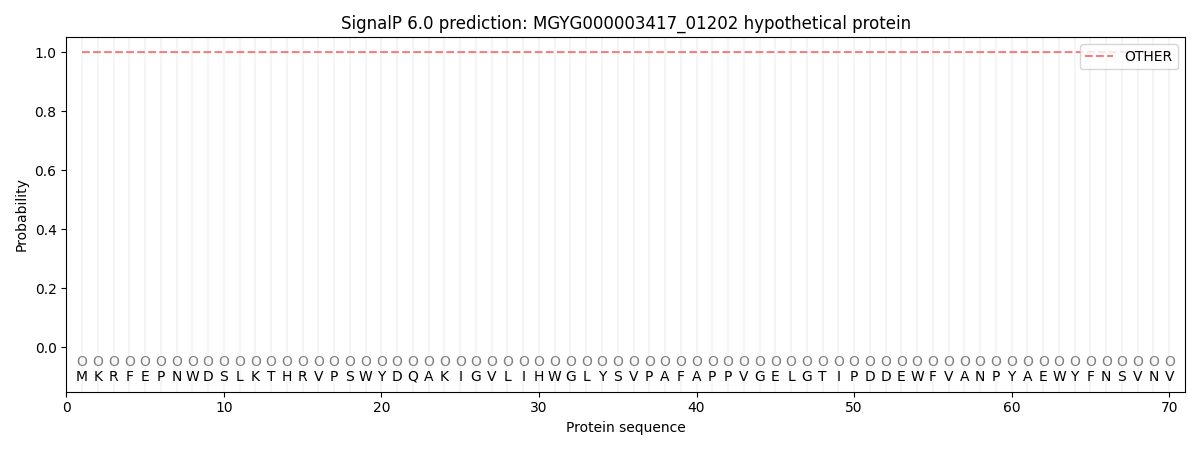
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999976 | 0.000055 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
