You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003351_02295
You are here: Home > Sequence: MGYG000003351_02295
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp900765785 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp900765785 | |||||||||||
| CAZyme ID | MGYG000003351_02295 | |||||||||||
| CAZy Family | GH97 | |||||||||||
| CAZyme Description | Glucan 1,4-alpha-glucosidase SusB | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 18411; End: 20420 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH97 | 13 | 666 | 1.6e-234 | 0.9920760697305864 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam10566 | Glyco_hydro_97 | 2.13e-147 | 276 | 565 | 2 | 277 | Glycoside hydrolase 97. This domain is the catalytic region of the bacterial glycosyl-hydrolase family 97. This central part of the GH97 family protein sequences represents a typical and complete (beta/alpha)8-barrel or catalytic TIM-barrel type domain. The N- and C-terminal parts of the sequences, mainly consisting of beta-strands, form two additional non-catalytic domains. In all known glycosidases with the (beta-alpha)8-barrel fold, the amino acid residues at the active site are located on the C-termini of the beta-strands. |
| pfam14508 | GH97_N | 1.05e-85 | 28 | 270 | 1 | 235 | Glycosyl-hydrolase 97 N-terminal. This N-terminal domain of glycosyl-hydrolase-97 contributes part of the active site pocket. It is also important for contact with the catalytic and C-terminal domains of the whole. |
| pfam14509 | GH97_C | 5.92e-44 | 569 | 667 | 1 | 97 | Glycosyl-hydrolase 97 C-terminal, oligomerization. Glycosyl-hydrolase-97 is made up of three tightly linked and highly conserved globular domains. The C-terminal domain is found to be necessary for oligomerization of the whole molecule in order to create the active-site pocket and the Ca++-binding site. |
| cd06563 | GH20_chitobiase-like | 0.001 | 288 | 372 | 3 | 111 | The chitobiase of Serratia marcescens is a beta-N-1,4-acetylhexosaminidase with a glycosyl hydrolase family 20 (GH20) domain that hydrolyzes the beta-1,4-glycosidic linkages in oligomers derived from chitin. Chitin is degraded by a two step process: i) a chitinase hydrolyzes the chitin to oligosaccharides and disaccharides such as di-N-acetyl-D-glucosamine and chitobiose, ii) chitobiase then further degrades these oligomers into monomers. This GH20 domain family includes an N-acetylglucosamidase (GlcNAcase A) from Pseudoalteromonas piscicida and an N-acetylhexosaminidase (SpHex) from Streptomyces plicatus. SpHex lacks the C-terminal PKD (polycystic kidney disease I)-like domain found in the chitobiases. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIU96187.1 | 0.0 | 1 | 669 | 1 | 669 |
| QMW87725.1 | 0.0 | 1 | 667 | 1 | 667 |
| AAO75790.1 | 0.0 | 1 | 667 | 1 | 667 |
| QNL38009.1 | 0.0 | 1 | 669 | 1 | 669 |
| QUU09059.1 | 0.0 | 1 | 669 | 1 | 669 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5HQ4_A | 1.36e-198 | 26 | 669 | 3 | 661 | AGlycoside Hydrolase Family 97 enzyme from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8],5HQA_A A Glycoside Hydrolase Family 97 enzyme in complex with Acarbose from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8] |
| 5HQC_A | 1.36e-198 | 26 | 669 | 3 | 661 | AGlycoside Hydrolase Family 97 enzyme R171K variant from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8] |
| 5HQB_A | 3.86e-198 | 26 | 669 | 3 | 661 | AGlycoside Hydrolase Family 97 enzyme (E480Q) in complex with Panose from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8] |
| 3WFA_A | 4.68e-166 | 25 | 665 | 4 | 702 | Catalyticrole of the calcium ion in GH97 inverting glycoside hydrolase [Bacteroides thetaiotaomicron],3WFA_B Catalytic role of the calcium ion in GH97 inverting glycoside hydrolase [Bacteroides thetaiotaomicron] |
| 2D73_A | 4.77e-166 | 20 | 665 | 19 | 722 | CrystalStructure Analysis of SusB [Bacteroides thetaiotaomicron VPI-5482],2D73_B Crystal Structure Analysis of SusB [Bacteroides thetaiotaomicron VPI-5482],2ZQ0_A Crystal structure of SusB complexed with acarbose [Bacteroides thetaiotaomicron],2ZQ0_B Crystal structure of SusB complexed with acarbose [Bacteroides thetaiotaomicron] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| G8JZS4 | 3.29e-166 | 2 | 665 | 5 | 722 | Glucan 1,4-alpha-glucosidase SusB OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=susB PE=1 SV=1 |
| Q8A6L0 | 1.05e-56 | 2 | 664 | 3 | 659 | Retaining alpha-galactosidase OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=BT_1871 PE=1 SV=1 |
| D7CFN7 | 1.48e-31 | 29 | 610 | 46 | 574 | Probable retaining alpha-galactosidase OS=Streptomyces bingchenggensis (strain BCW-1) OX=749414 GN=SBI_01652 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
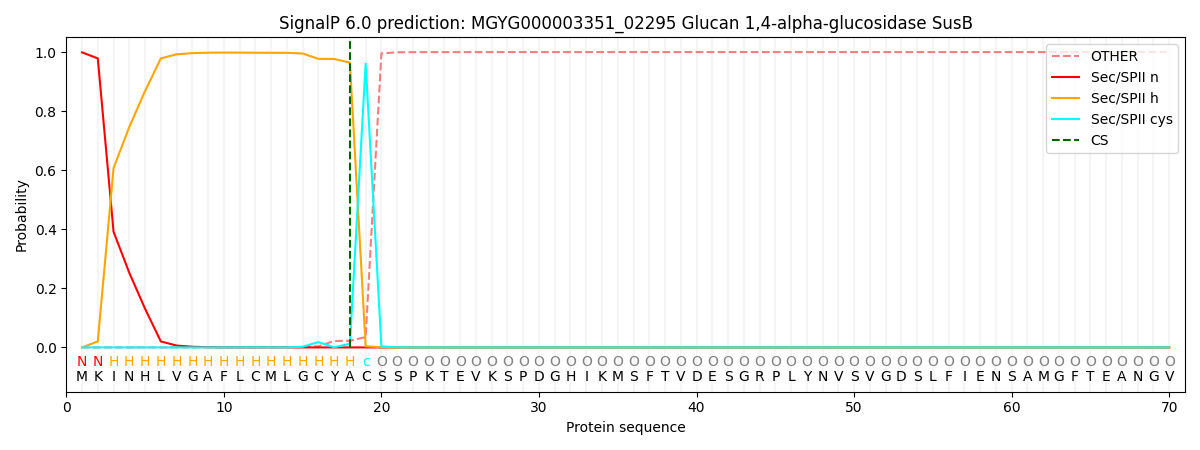
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000006 | 0.000930 | 0.999127 | 0.000000 | 0.000001 | 0.000000 |
