You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002846_01010
You are here: Home > Sequence: MGYG000002846_01010
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
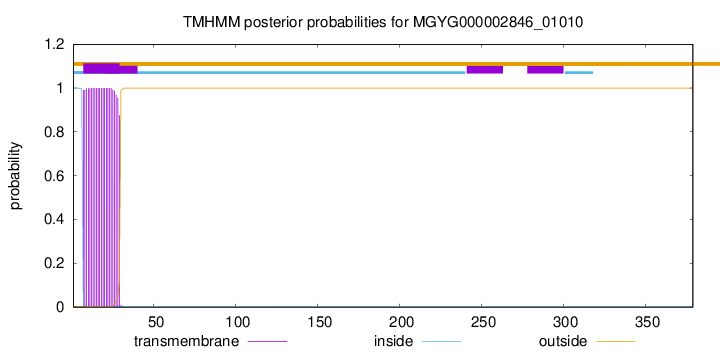
TMHMM annotations
Basic Information help
| Species | Desulfovibrio sp900540515 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Desulfobacterota; Desulfovibrionia; Desulfovibrionales; Desulfovibrionaceae; Desulfovibrio; Desulfovibrio sp900540515 | |||||||||||
| CAZyme ID | MGYG000002846_01010 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase C | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 23993; End: 25132 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 210 | 371 | 3e-22 | 0.8740740740740741 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16893 | LT_MltC_MltE | 5.39e-68 | 214 | 372 | 3 | 161 | membrane-bound lytic murein transglycosylases MltC and MltE, and similar proteins. MltC and MltE are periplasmic, outer membrane attached lytic transglycosylases (LTs), which cleave beta-1,4-glycosidic bonds joining N-acetylmuramic acid and N-acetylglucosamine in the cell wall peptidoglycan, yielding 1,6-anhydromuropeptides. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda |
| PRK11671 | mltC | 2.36e-38 | 143 | 375 | 127 | 356 | membrane-bound lytic murein transglycosylase MltC. |
| cd16896 | LT_Slt70-like | 2.61e-35 | 209 | 370 | 3 | 142 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd13401 | Slt70-like | 1.17e-30 | 208 | 368 | 4 | 142 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| PRK15470 | emtA | 6.03e-26 | 206 | 370 | 35 | 198 | membrane-bound lytic murein transglycosylase EmtA. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AMD88640.1 | 6.69e-234 | 1 | 379 | 1 | 379 |
| QCC84560.1 | 4.47e-148 | 3 | 379 | 3 | 392 |
| VZH33299.1 | 2.52e-146 | 5 | 378 | 5 | 356 |
| ATD80276.1 | 2.70e-146 | 3 | 378 | 3 | 358 |
| SPD35752.1 | 2.70e-146 | 3 | 378 | 3 | 358 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4C5F_A | 1.06e-28 | 141 | 375 | 107 | 338 | Structureof Lytic Transglycosylase MltC from Escherichia coli at 2.3 A resolution. [Escherichia coli],4C5F_B Structure of Lytic Transglycosylase MltC from Escherichia coli at 2.3 A resolution. [Escherichia coli] |
| 4CFO_A | 2.79e-28 | 141 | 375 | 107 | 338 | Structureof Lytic Transglycosylase MltC from Escherichia coli in complex with tetrasaccharide at 2.9 A resolution. [Escherichia coli],4CFO_B Structure of Lytic Transglycosylase MltC from Escherichia coli in complex with tetrasaccharide at 2.9 A resolution. [Escherichia coli],4CFP_A Crystal structure of MltC in complex with tetrasaccharide at 2.15 A resolution [Escherichia coli],4CFP_B Crystal structure of MltC in complex with tetrasaccharide at 2.15 A resolution [Escherichia coli],4CHX_A Crystal structure of MltC in complex with disaccharide pentapeptide DHl89 [Escherichia coli],4CHX_B Crystal structure of MltC in complex with disaccharide pentapeptide DHl89 [Escherichia coli] |
| 2Y8P_A | 9.46e-20 | 206 | 368 | 18 | 179 | CrystalStructure of an Outer Membrane-Anchored Endolytic Peptidoglycan Lytic Transglycosylase (MltE) from Escherichia coli [Escherichia coli K-12],2Y8P_B Crystal Structure of an Outer Membrane-Anchored Endolytic Peptidoglycan Lytic Transglycosylase (MltE) from Escherichia coli [Escherichia coli K-12] |
| 3T36_A | 1.15e-19 | 206 | 368 | 35 | 196 | Crystalstructure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_B Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_C Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_D Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_E Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],4HJV_A Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_B Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_C Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_D Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_E Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12] |
| 6GI4_B | 1.30e-19 | 206 | 368 | 18 | 179 | Structureof Lytic Transglycosylase MltE mutant S75A from E.coli [Escherichia coli] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A8GJ50 | 6.36e-32 | 143 | 375 | 126 | 355 | Membrane-bound lytic murein transglycosylase C OS=Serratia proteamaculans (strain 568) OX=399741 GN=mltC PE=3 SV=1 |
| C5BCE9 | 2.33e-30 | 146 | 379 | 129 | 359 | Membrane-bound lytic murein transglycosylase C OS=Edwardsiella ictaluri (strain 93-146) OX=634503 GN=mltC PE=3 SV=1 |
| A4WEA0 | 1.60e-29 | 143 | 375 | 126 | 355 | Membrane-bound lytic murein transglycosylase C OS=Enterobacter sp. (strain 638) OX=399742 GN=mltC PE=3 SV=2 |
| B2K0V2 | 3.06e-29 | 143 | 375 | 126 | 355 | Membrane-bound lytic murein transglycosylase C OS=Yersinia pseudotuberculosis serotype IB (strain PB1/+) OX=502801 GN=mltC PE=3 SV=1 |
| Q1CB94 | 3.06e-29 | 143 | 375 | 126 | 355 | Membrane-bound lytic murein transglycosylase C OS=Yersinia pestis bv. Antiqua (strain Antiqua) OX=360102 GN=mltC PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
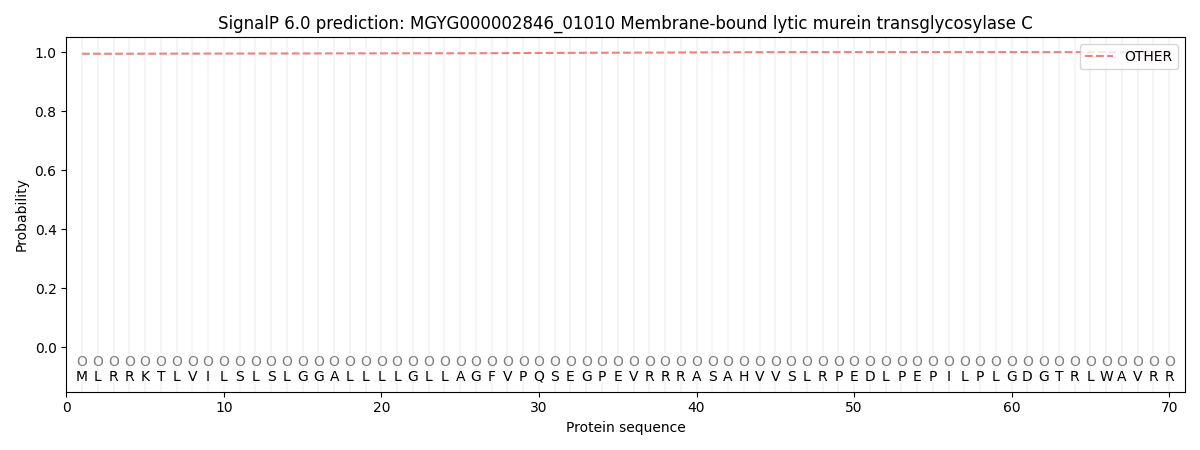
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.994590 | 0.004134 | 0.000585 | 0.000037 | 0.000020 | 0.000664 |