You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002795_01105
You are here: Home > Sequence: MGYG000002795_01105
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
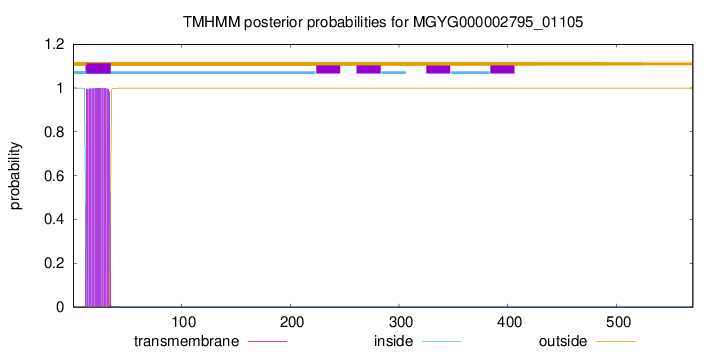
TMHMM annotations
Basic Information help
| Species | CAG-110 sp900548795 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; CAG-110; CAG-110 sp900548795 | |||||||||||
| CAZyme ID | MGYG000002795_01105 | |||||||||||
| CAZy Family | GT4 | |||||||||||
| CAZyme Description | Glycosyltransferase Gtf1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1357; End: 3069 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd03808 | GT4_CapM-like | 1.48e-117 | 207 | 560 | 1 | 358 | capsular polysaccharide biosynthesis glycosyltransferase CapM and similar proteins. This family is most closely related to the GT4 family of glycosyltransferases. CapM in Staphylococcus aureus is required for the synthesis of type 1 capsular polysaccharides. |
| pfam02397 | Bac_transf | 4.66e-80 | 7 | 186 | 1 | 181 | Bacterial sugar transferase. This Pfam family represents a conserved region from a number of different bacterial sugar transferases, involved in diverse biosynthesis pathways. |
| COG2148 | WcaJ | 3.45e-72 | 1 | 186 | 35 | 221 | Sugar transferase involved in LPS biosynthesis (colanic, teichoic acid) [Cell wall/membrane/envelope biogenesis]. |
| TIGR03025 | EPS_sugtrans | 3.22e-59 | 6 | 181 | 257 | 435 | exopolysaccharide biosynthesis polyprenyl glycosylphosphotransferase. Members of this family are generally found near other genes involved in the biosynthesis of a variety of exopolysaccharides. These proteins consist of two fused domains, an N-terminal hydrophobic domain of generally low conservation and a highly conserved C-terminal sugar transferase domain (pfam02397). Characterized and partially characterized members of this subfamily include Salmonella WbaP (originally RfbP), E. coli WcaJ, Methylobacillus EpsB, Xanthomonas GumD, Vibrio CpsA, Erwinia AmsG, Group B Streptococcus CpsE (originally CpsD), and Streptococcus suis Cps2E. Each of these is believed to act in transferring the sugar from, for instance, UDP-glucose or UDP-galactose, to a lipid carrier such as undecaprenyl phosphate as the first (priming) step in the synthesis of an oligosaccharide "block". This function is encoded in the C-terminal domain. The liposaccharide is believed to be subsequently transferred through a "flippase" function from the cytoplasmic to the periplasmic face of the inner membrane by the N-terminal domain. Certain closely related transferase enzymes, such as Sinorhizobium ExoY and Lactococcus EpsD, lack the N-terminal domain and are not found by this model. |
| cd03801 | GT4_PimA-like | 2.12e-49 | 224 | 563 | 22 | 365 | phosphatidyl-myo-inositol mannosyltransferase. This family is most closely related to the GT4 family of glycosyltransferases and named after PimA in Propionibacterium freudenreichii, which is involved in the biosynthesis of phosphatidyl-myo-inositol mannosides (PIM) which are early precursors in the biosynthesis of lipomannans (LM) and lipoarabinomannans (LAM), and catalyzes the addition of a mannosyl residue from GDP-D-mannose (GDP-Man) to the position 2 of the carrier lipid phosphatidyl-myo-inositol (PI) to generate a phosphatidyl-myo-inositol bearing an alpha-1,2-linked mannose residue (PIM1). Glycosyltransferases catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. The acceptor molecule can be a lipid, a protein, a heterocyclic compound, or another carbohydrate residue. This group of glycosyltransferases is most closely related to the previously defined glycosyltransferase family 1 (GT1). The members of this family may transfer UDP, ADP, GDP, or CMP linked sugars. The diverse enzymatic activities among members of this family reflect a wide range of biological functions. The protein structure available for this family has the GTB topology, one of the two protein topologies observed for nucleotide-sugar-dependent glycosyltransferases. GTB proteins have distinct N- and C- terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. The members of this family are found mainly in certain bacteria and archaea. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ANU78758.2 | 2.21e-158 | 204 | 556 | 12 | 368 |
| QQQ95319.1 | 2.21e-158 | 204 | 556 | 12 | 368 |
| ASU30524.1 | 2.21e-158 | 204 | 556 | 12 | 368 |
| QJU16507.1 | 2.21e-158 | 204 | 556 | 12 | 368 |
| ALP95394.1 | 5.90e-152 | 207 | 563 | 2 | 357 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5W7L_A | 1.44e-30 | 1 | 200 | 5 | 200 | Structureof Campylobacter concisus PglC I57M/Q175M variant [Campylobacter concisus 13826],5W7L_B Structure of Campylobacter concisus PglC I57M/Q175M variant [Campylobacter concisus 13826] |
| 7MI0_A | 4.77e-06 | 195 | 525 | 32 | 350 | ChainA, Glycosyltransferase [Rickettsia africae ESF-5] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0P9D0 | 3.65e-30 | 1 | 194 | 1 | 189 | Undecaprenyl phosphate N,N'-diacetylbacillosamine 1-phosphate transferase OS=Campylobacter jejuni subsp. jejuni serotype O:2 (strain ATCC 700819 / NCTC 11168) OX=192222 GN=pglC PE=1 SV=1 |
| Q0P9C9 | 4.10e-30 | 218 | 528 | 13 | 336 | N,N'-diacetylbacillosaminyl-diphospho-undecaprenol alpha-1,3-N-acetylgalactosaminyltransferase OS=Campylobacter jejuni subsp. jejuni serotype O:2 (strain ATCC 700819 / NCTC 11168) OX=192222 GN=pglA PE=1 SV=1 |
| P71062 | 4.86e-29 | 6 | 190 | 3 | 183 | Uncharacterized sugar transferase EpsL OS=Bacillus subtilis (strain 168) OX=224308 GN=epsL PE=2 SV=1 |
| P71241 | 2.51e-27 | 6 | 180 | 273 | 453 | UDP-glucose:undecaprenyl-phosphate glucose-1-phosphate transferase OS=Escherichia coli (strain K12) OX=83333 GN=wcaJ PE=1 SV=2 |
| P14186 | 1.01e-26 | 7 | 187 | 29 | 224 | Exopolysaccharide production protein ExoY OS=Sinorhizobium fredii (strain NBRC 101917 / NGR234) OX=394 GN=exoY PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
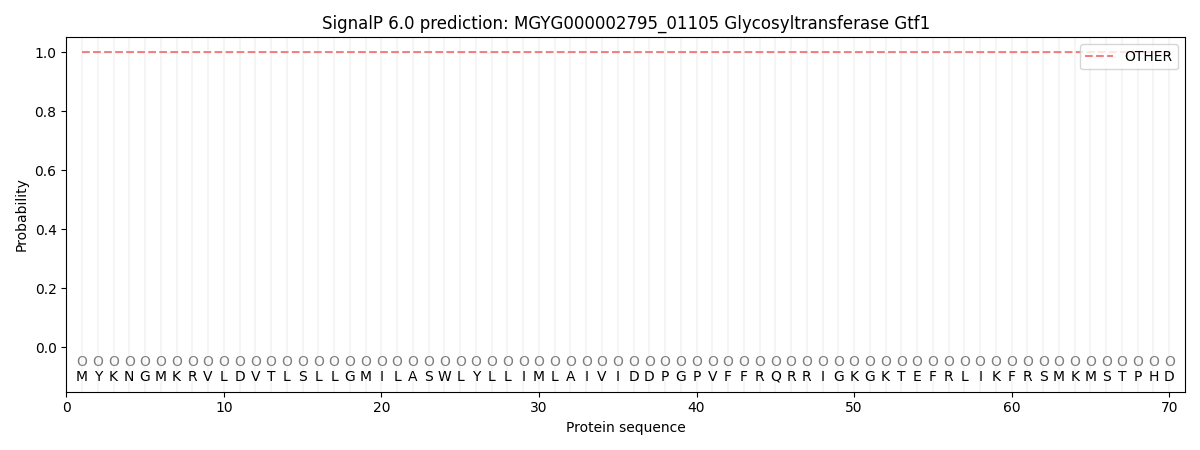
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000071 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |