You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002535_01657
You are here: Home > Sequence: MGYG000002535_01657
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Citrobacter gillenii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Citrobacter; Citrobacter gillenii | |||||||||||
| CAZyme ID | MGYG000002535_01657 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Endo-type membrane-bound lytic murein transglycosylase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1732685; End: 1733296 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 38 | 196 | 8.3e-17 | 0.837037037037037 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK15470 | emtA | 2.86e-157 | 1 | 203 | 1 | 203 | membrane-bound lytic murein transglycosylase EmtA. |
| cd16893 | LT_MltC_MltE | 2.61e-82 | 41 | 199 | 1 | 160 | membrane-bound lytic murein transglycosylases MltC and MltE, and similar proteins. MltC and MltE are periplasmic, outer membrane attached lytic transglycosylases (LTs), which cleave beta-1,4-glycosidic bonds joining N-acetylmuramic acid and N-acetylglucosamine in the cell wall peptidoglycan, yielding 1,6-anhydromuropeptides. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda |
| PRK11671 | mltC | 5.13e-58 | 34 | 199 | 187 | 352 | membrane-bound lytic murein transglycosylase MltC. |
| pfam01464 | SLT | 4.48e-32 | 45 | 168 | 3 | 114 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| COG0741 | MltE | 8.23e-29 | 1 | 201 | 93 | 296 | Soluble lytic murein transglycosylase and related regulatory proteins (some contain LysM/invasin domains) [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QMR43876.1 | 1.41e-147 | 1 | 203 | 1 | 203 |
| ATF49186.1 | 1.41e-147 | 1 | 203 | 1 | 203 |
| QCA18084.1 | 1.41e-147 | 1 | 203 | 1 | 203 |
| QBX80114.1 | 5.76e-147 | 1 | 203 | 1 | 203 |
| QNM21544.1 | 1.16e-146 | 1 | 203 | 1 | 203 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3T36_A | 7.38e-124 | 17 | 203 | 17 | 203 | Crystalstructure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_B Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_C Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_D Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],3T36_E Crystal structure of lytic transglycosylase MltE from Eschericha coli [Escherichia coli K-12],4HJV_A Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_B Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_C Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_D Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12],4HJV_E Crystal structure of E. coli MltE with bound bulgecin and murodipeptide [Escherichia coli K-12] |
| 4HJY_A | 2.35e-123 | 17 | 203 | 20 | 206 | 2.4A Crystal structure of E. coli MltE-E64Q with bound chitopentaose [Escherichia coli K-12],4HJY_B 2.4 A Crystal structure of E. coli MltE-E64Q with bound chitopentaose [Escherichia coli K-12],4HJZ_A 1.9 A Crystal structure of E. coli MltE-E64Q with bound chitopentaose [Escherichia coli K-12],4HJZ_B 1.9 A Crystal structure of E. coli MltE-E64Q with bound chitopentaose [Escherichia coli K-12] |
| 2Y8P_A | 8.90e-123 | 19 | 203 | 2 | 186 | CrystalStructure of an Outer Membrane-Anchored Endolytic Peptidoglycan Lytic Transglycosylase (MltE) from Escherichia coli [Escherichia coli K-12],2Y8P_B Crystal Structure of an Outer Membrane-Anchored Endolytic Peptidoglycan Lytic Transglycosylase (MltE) from Escherichia coli [Escherichia coli K-12] |
| 6GHY_A | 2.55e-122 | 19 | 203 | 2 | 186 | Structureof Lytic Transglycosylase MltE inactive mutant E64Q from E.coli [Escherichia coli K-12],6GHY_B Structure of Lytic Transglycosylase MltE inactive mutant E64Q from E.coli [Escherichia coli K-12] |
| 6GI3_B | 2.55e-122 | 19 | 203 | 2 | 186 | Structureof Lytic Transglycosylase MltE mutant S73A from E.coli [Escherichia coli] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q57NL3 | 4.65e-138 | 1 | 203 | 1 | 203 | Endo-type membrane-bound lytic murein transglycosylase A OS=Salmonella choleraesuis (strain SC-B67) OX=321314 GN=emtA PE=3 SV=1 |
| B5R901 | 4.65e-138 | 1 | 203 | 1 | 203 | Endo-type membrane-bound lytic murein transglycosylase A OS=Salmonella gallinarum (strain 287/91 / NCTC 13346) OX=550538 GN=emtA PE=3 SV=1 |
| Q5PN08 | 4.65e-138 | 1 | 203 | 1 | 203 | Endo-type membrane-bound lytic murein transglycosylase A OS=Salmonella paratyphi A (strain ATCC 9150 / SARB42) OX=295319 GN=emtA PE=3 SV=1 |
| Q7CQE4 | 4.65e-138 | 1 | 203 | 1 | 203 | Endo-type membrane-bound lytic murein transglycosylase A OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=emtA PE=3 SV=1 |
| C0Q334 | 4.65e-138 | 1 | 203 | 1 | 203 | Endo-type membrane-bound lytic murein transglycosylase A OS=Salmonella paratyphi C (strain RKS4594) OX=476213 GN=emtA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
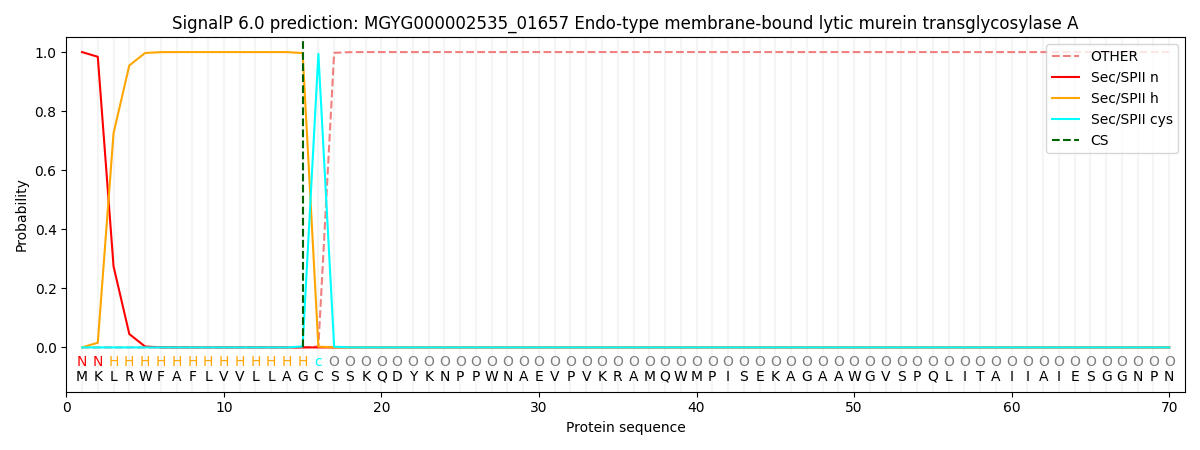
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000056 | 0.000000 | 0.000000 | 0.000000 |
