You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002490_03907
You are here: Home > Sequence: MGYG000002490_03907
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
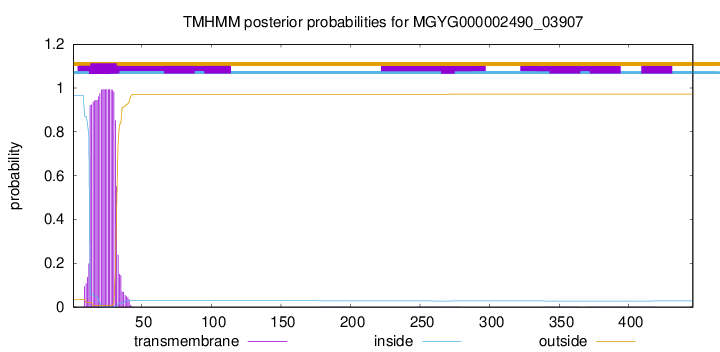
TMHMM annotations
Basic Information help
| Species | Paenibacillus_B thiaminolyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_B; Paenibacillus_B thiaminolyticus | |||||||||||
| CAZyme ID | MGYG000002490_03907 | |||||||||||
| CAZy Family | AA10 | |||||||||||
| CAZyme Description | GlcNAc-binding protein A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 258248; End: 259588 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA10 | 38 | 202 | 2.8e-54 | 0.9943820224719101 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK13211 | PRK13211 | 1.18e-101 | 15 | 212 | 5 | 202 | N-acetylglucosamine-binding protein GbpA. |
| COG3397 | COG3397 | 5.28e-78 | 20 | 294 | 12 | 296 | Predicted carbohydrate-binding protein, contains CBM5 and CBM33 domains [General function prediction only]. |
| cd21177 | LPMO_AA10 | 1.59e-67 | 38 | 202 | 1 | 180 | lytic polysaccharide monooxygenase (LPMO) auxiliary activity family 10 (AA10). AA10 proteins are copper-dependent lytic polysaccharide monooxygenases (LPMOs), which may act on chitin or cellulose. The family used to be called CBM33. Activities in this family include lytic cellulose monooxygenase (C1-hydroxylating) (EC 1.14.99.54), lytic cellulose monooxygenase (C4-dehydrogenating) (EC 1.14.99.56), lytic chitin monooxygenase (EC 1.14.99.53), and lytic xylan monooxygenase/xylan oxidase (glycosidic bond-cleaving) (EC 1.14.99.-). Also included are viral chitin-binding glycoproteins such as fusolin and spheroidin-like proteins. |
| pfam03067 | LPMO_10 | 4.55e-55 | 38 | 201 | 1 | 186 | Lytic polysaccharide mono-oxygenase, cellulose-degrading. This domain is found associated with a wide variety of cellulose binding domains. This is a family of two very closely related proteins that together act as both a C1- and a C4-oxidising lytic polysaccharide mono-oxygenase, degrading cellulose. This domain is also found in baculoviral spheroidins and spindolins, protein of unknown function. |
| cd00063 | FN3 | 2.88e-15 | 214 | 300 | 1 | 93 | Fibronectin type 3 domain; One of three types of internal repeats found in the plasma protein fibronectin. Its tenth fibronectin type III repeat contains an RGD cell recognition sequence in a flexible loop between 2 strands. Approximately 2% of all animal proteins contain the FN3 repeat; including extracellular and intracellular proteins, membrane spanning cytokine receptors, growth hormone receptors, tyrosine phosphatase receptors, and adhesion molecules. FN3-like domains are also found in bacterial glycosyl hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDM42328.1 | 0.0 | 1 | 446 | 3 | 448 |
| SYX83687.1 | 2.46e-252 | 12 | 446 | 14 | 448 |
| QAV18977.1 | 3.44e-203 | 8 | 446 | 8 | 450 |
| QAV18986.1 | 2.61e-202 | 1 | 446 | 1 | 448 |
| QDM42769.1 | 3.74e-195 | 6 | 446 | 6 | 449 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2BEM_A | 1.45e-61 | 38 | 202 | 1 | 167 | Crystalstructure of the Serratia marcescens chitin-binding protein CBP21 [Serratia marcescens],2BEM_B Crystal structure of the Serratia marcescens chitin-binding protein CBP21 [Serratia marcescens],2BEM_C Crystal structure of the Serratia marcescens chitin-binding protein CBP21 [Serratia marcescens],2LHS_A Structure of the chitin binding protein 21 (CBP21) [Serratia marcescens] |
| 2BEN_A | 3.17e-60 | 38 | 202 | 1 | 167 | Crystalstructure of the Serratia marcescens chitin-binding protein CBP21 Y54A mutant. [Serratia marcescens],2BEN_B Crystal structure of the Serratia marcescens chitin-binding protein CBP21 Y54A mutant. [Serratia marcescens] |
| 5WSZ_A | 4.75e-59 | 38 | 202 | 1 | 165 | Crystalstructure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki],5WSZ_B Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki],5WSZ_C Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki],5WSZ_D Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki] |
| 2XWX_A | 2.78e-57 | 38 | 211 | 1 | 186 | Vibriocholerae colonization factor GbpA crystal structure [Vibrio cholerae],2XWX_B Vibrio cholerae colonization factor GbpA crystal structure [Vibrio cholerae] |
| 6T5Z_A | 1.45e-55 | 38 | 200 | 1 | 168 | Crystalstructure of an AA10 LPMO from Photorhabdus luminescens [Photorhabdus laumondii subsp. laumondii TTO1],6T5Z_B Crystal structure of an AA10 LPMO from Photorhabdus luminescens [Photorhabdus laumondii subsp. laumondii TTO1],6T5Z_C Crystal structure of an AA10 LPMO from Photorhabdus luminescens [Photorhabdus laumondii subsp. laumondii TTO1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P20533 | 1.30e-72 | 166 | 446 | 408 | 699 | Chitinase A1 OS=Niallia circulans OX=1397 GN=chiA1 PE=1 SV=1 |
| Q8D7V4 | 5.53e-61 | 24 | 211 | 10 | 208 | GlcNAc-binding protein A OS=Vibrio vulnificus (strain CMCP6) OX=216895 GN=gbpA PE=3 SV=1 |
| Q7MEW9 | 5.53e-61 | 24 | 211 | 10 | 208 | GlcNAc-binding protein A OS=Vibrio vulnificus (strain YJ016) OX=196600 GN=gbpA PE=3 SV=1 |
| Q8EHY2 | 8.63e-61 | 22 | 209 | 12 | 201 | GlcNAc-binding protein A OS=Shewanella oneidensis (strain MR-1) OX=211586 GN=gbpA PE=3 SV=2 |
| Q5E183 | 2.60e-59 | 17 | 212 | 4 | 214 | GlcNAc-binding protein A OS=Aliivibrio fischeri (strain ATCC 700601 / ES114) OX=312309 GN=gbpA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
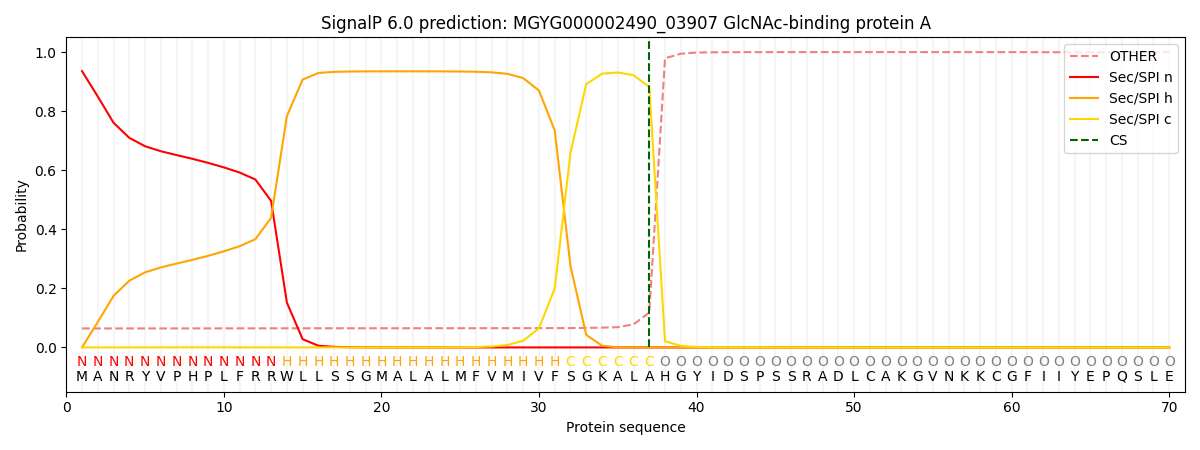
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.071200 | 0.926875 | 0.000626 | 0.000597 | 0.000337 | 0.000322 |