You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002349_00754
You are here: Home > Sequence: MGYG000002349_00754
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Streptococcus agalactiae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus agalactiae | |||||||||||
| CAZyme ID | MGYG000002349_00754 | |||||||||||
| CAZy Family | GT1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10255; End: 10704 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam08660 | Alg14 | 1.57e-25 | 3 | 146 | 1 | 169 | Oligosaccharide biosynthesis protein Alg14 like. Alg14 is involved dolichol-linked oligosaccharide biosynthesis and anchors the catalytic subunit Alg13 to the ER membrane. |
| COG0707 | MurG | 1.43e-11 | 1 | 142 | 1 | 154 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
| cd03785 | GT28_MurG | 2.37e-05 | 6 | 142 | 6 | 152 | undecaprenyldiphospho-muramoylpentapeptide beta-N-acetylglucosaminyltransferase. MurG (EC 2.4.1.227) is an N-acetylglucosaminyltransferase, the last enzyme involved in the intracellular phase of peptidoglycan biosynthesis. It transfers N-acetyl-D-glucosamine (GlcNAc) from UDP-GlcNAc to the C4 hydroxyl of a lipid-linked N-acetylmuramoyl pentapeptide (NAM). The resulting disaccharide is then transported across the cell membrane, where it is polymerized into NAG-NAM cell-wall repeat structure. MurG belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains, each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
| cd03820 | GT4_AmsD-like | 0.002 | 50 | 144 | 45 | 158 | amylovoran biosynthesis glycosyltransferase AmsD and similar proteins. This family is most closely related to the GT4 family of glycosyltransferases. AmSD in Erwinia amylovora has been shown to be involved in the biosynthesis of amylovoran, the acidic exopolysaccharide acting as a virulence factor. This enzyme may be responsible for the formation of galactose alpha-1,6 linkages in amylovoran. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AIK75892.1 | 1.47e-109 | 1 | 149 | 1 | 149 |
| ABG38180.1 | 1.47e-109 | 1 | 149 | 1 | 149 |
| SIO74009.1 | 1.47e-109 | 1 | 149 | 1 | 149 |
| QHO95688.1 | 1.47e-109 | 1 | 149 | 1 | 149 |
| ASA88077.1 | 1.47e-109 | 1 | 149 | 1 | 149 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q96F25 | 2.22e-11 | 53 | 148 | 106 | 215 | UDP-N-acetylglucosamine transferase subunit ALG14 homolog OS=Homo sapiens OX=9606 GN=ALG14 PE=1 SV=1 |
| Q6AY85 | 2.20e-10 | 68 | 148 | 129 | 215 | UDP-N-acetylglucosamine transferase subunit ALG14 homolog OS=Rattus norvegicus OX=10116 GN=Alg14 PE=2 SV=1 |
| Q5A5N6 | 3.16e-10 | 2 | 146 | 49 | 216 | UDP-N-acetylglucosamine transferase subunit ALG14 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=ALG14 PE=3 SV=1 |
| Q9D081 | 1.58e-09 | 68 | 148 | 130 | 216 | UDP-N-acetylglucosamine transferase subunit ALG14 homolog OS=Mus musculus OX=10090 GN=Alg14 PE=1 SV=1 |
| Q6CJG3 | 2.61e-09 | 40 | 146 | 108 | 233 | UDP-N-acetylglucosamine transferase subunit ALG14 OS=Kluyveromyces lactis (strain ATCC 8585 / CBS 2359 / DSM 70799 / NBRC 1267 / NRRL Y-1140 / WM37) OX=284590 GN=ALG14 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
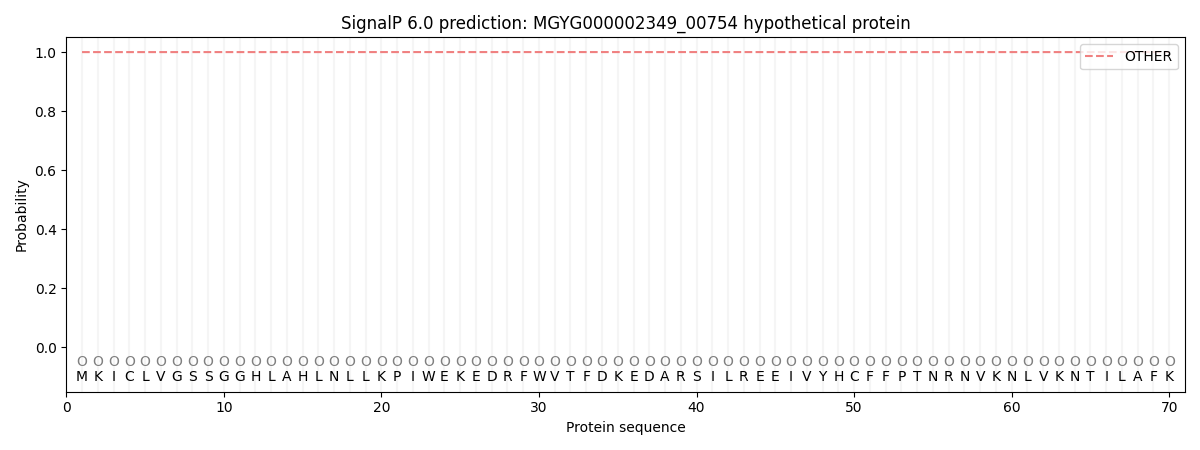
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000051 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
