You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002273_02457
You are here: Home > Sequence: MGYG000002273_02457
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides stercorirosoris | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides stercorirosoris | |||||||||||
| CAZyme ID | MGYG000002273_02457 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 37638; End: 39008 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH43 | 49 | 283 | 5.3e-72 | 0.9878048780487805 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd18821 | GH43_Pc3Gal43A-like | 8.95e-145 | 41 | 300 | 1 | 262 | Glycosyl hydrolase family 43 protein such as Phanerochaete chrysosporium exo-beta-1,3-galactanase (Pc1, 3Gal43A, 1,3Gal43A). This glycosyl hydrolase family 43 (GH43) subgroup includes characterized enzymes with exo-beta-1,3-galactanase (EC 3.2.1.145, also known as galactan 1,3-beta-galactosidase) activity such as Phanerochaete chrysosporium 1,3Gal43A (Pc1, 3Gal43A), Fusarium oxysporum 12S Fo/1 (3Gal), and Streptomyces sp. 19(2012) SGalase1 and SGalase2. It belongs to the GH43_CtGH43 subgroup of the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43_CtGH43 includes proteins such as Clostridium thermocellum exo-beta-1,3-galactanase (Ct1,3Gal43A or CtGH43) which is comprised of the GH43 domain, a CBM13 domain, and a dockerin domain, exhibits an unusual ability to hydrolyze beta-1,3-galactan in the presence of a beta-1,6 linked branch, and is missing an essential acidic residue suggesting a mechanism by which it bypasses beta-1,6 linked branches in the substrate. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd08985 | GH43_CtGH43-like | 1.52e-75 | 41 | 298 | 1 | 271 | Glycosyl hydrolase family 43 protein such as Clostridium thermocellum exo-beta-1,3-galactanase CtGH43 and Ruminococcus champanellensis arabinanase Ara43A. This glycosyl hydrolase family 43 (GH43) subgroup includes characterized enzymes with exo-beta-1,3-galactanase (EC 3.2.1.145, also known as galactan 1,3-beta-galactosidase) activity such as Clostridium thermocellum (Ct1,3Gal43A or CtGH43) and Phanerochaete chrysosporium 1,3Gal43A (Pc1, 3Gal43A), and arabinanase (EC 3.2.1.99) activity such as Ruminococcus champanellensis Ara43A. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd18822 | GH43_CtGH43-like | 2.79e-71 | 41 | 299 | 1 | 265 | Glycosyl hydrolase family 43 protein such as Clostridium thermocellum exo-beta-1,3-galactanase (Ct1,3Gal43A or CtGH43). This glycosyl hydrolase family 43 (GH43) subgroup includes characterized enzymes with exo-beta-1,3-galactanase (EC 3.2.1.145, also known as galactan 1,3-beta-galactosidase) activity such as Clostridium thermocellum exo-beta-1,3-galactanase (Ct1,3Gal43A or CtGH43), Streptomyces avermitilis MA-4680 = NBRC 14893 (Sa1,3Gal43A;SAV2109) (1,3Gal43A), and Ruminiclostridium thermocellum ATCC 27405 (Ct1,3Gal43A;CtGH43;Cthe_0661) (1,3Gal43A). It belongs to the GH43_CtGH43 subgroup of the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43_CtGH43 includes proteins such as Clostridium thermocellum exo-beta-1,3-galactanase (Ct1,3Gal43A or CtGH43) which is comprised of the GH43 domain, a CBM13 domain, and a dockerin domain, exhibits an unusual ability to hydrolyze beta-1,3-galactan in the presence of a beta-1,6 linked branch, and is missing an essential acidic residue suggesting a mechanism by which it bypasses beta-1,6 linked branches in the substrate. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd18825 | GH43_CtGH43-like | 1.31e-70 | 41 | 300 | 1 | 285 | Glycosyl hydrolase family 43 protein similar to Clostridium thermocellum exo-beta-1,3-galactanase CtGH43 and Ruminococcus champanellensis arabinanase Ara43A. This uncharacterized glycosyl hydrolase family 43 (GH43) subgroup belongs to a subgroup which includes characterized enzymes with exo-beta-1,3-galactanase (EC 3.2.1.145, also known as galactan 1,3-beta-galactosidase) activity such as Clostridium thermocellum (Ct1,3Gal43A or CtGH43) and Phanerochaete chrysosporium 1,3Gal43A (Pc1, 3Gal43A), and arabinanase (EC 3.2.1.99) activity such as Ruminococcus champanellensis Ara43A. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd18826 | GH43_CtGH43-like | 1.75e-53 | 41 | 299 | 1 | 268 | Glycosyl hydrolase family 43 protein similar to Clostridium thermocellum exo-beta-1,3-galactanase CtGH43 and Ruminococcus champanellensis arabinanase Ara43A. This uncharacterized glycosyl hydrolase family 43 (GH43) subgroup belongs to a subgroup which includes characterized enzymes with exo-beta-1,3-galactanase (EC 3.2.1.145, also known as galactan 1,3-beta-galactosidase) activity such as Clostridium thermocellum (Ct1,3Gal43A or CtGH43) and Phanerochaete chrysosporium 1,3Gal43A (Pc1, 3Gal43A), and arabinanase (EC 3.2.1.99) activity such as Ruminococcus champanellensis Ara43A. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDO71467.1 | 0.0 | 6 | 456 | 6 | 456 |
| ALJ58889.1 | 1.89e-313 | 1 | 456 | 9 | 464 |
| QUT89994.1 | 2.68e-313 | 1 | 456 | 9 | 464 |
| SCV06763.1 | 4.13e-287 | 1 | 453 | 6 | 459 |
| ALJ47356.1 | 4.13e-287 | 1 | 453 | 6 | 459 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3NQH_A | 8.13e-239 | 25 | 453 | 6 | 438 | Crystalstructure of a glycosyl hydrolase (BT_2959) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.11 A resolution [Bacteroides thetaiotaomicron VPI-5482] |
| 5FLW_A | 6.82e-58 | 35 | 300 | 12 | 285 | Crystalstructure of putative exo-beta-1,3-galactanase from Bifidobacterium bifidum s17 [Bifidobacterium bifidum S17],5FLW_B Crystal structure of putative exo-beta-1,3-galactanase from Bifidobacterium bifidum s17 [Bifidobacterium bifidum S17] |
| 7BYS_A | 1.13e-48 | 27 | 453 | 3 | 424 | ChainA, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium],7BYS_B Chain B, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium],7BYT_A Chain A, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium] |
| 7BYV_A | 3.13e-48 | 27 | 453 | 4 | 425 | ChainA, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium] |
| 7BYX_A | 8.27e-48 | 27 | 453 | 3 | 424 | ChainA, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium],7BYX_B Chain B, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium],7BYX_C Chain C, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium],7BYX_D Chain D, Galactan 1,3-beta-galactosidase [Phanerodontia chrysosporium] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as LIPO
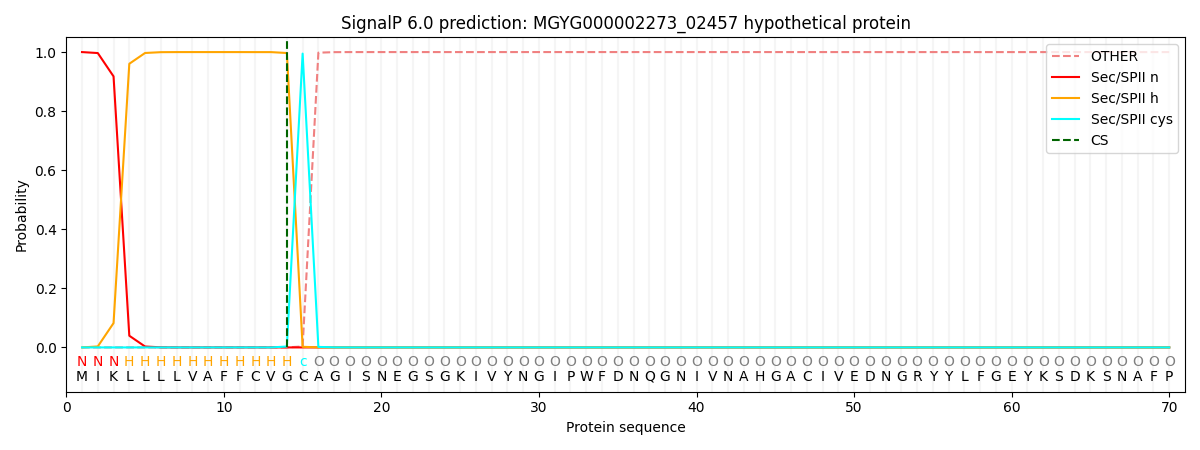
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000001 | 1.000069 | 0.000000 | 0.000000 | 0.000000 |
