You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001963_00079
You are here: Home > Sequence: MGYG000001963_00079
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola sp000434735 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola sp000434735 | |||||||||||
| CAZyme ID | MGYG000001963_00079 | |||||||||||
| CAZy Family | GH63 | |||||||||||
| CAZyme Description | Glucosidase YgjK | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 101077; End: 103077 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH63 | 199 | 663 | 2e-107 | 0.5631578947368421 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10137 | PRK10137 | 2.47e-154 | 30 | 657 | 29 | 774 | alpha-glucosidase; Provisional |
| COG3408 | GDB1 | 2.33e-24 | 306 | 665 | 268 | 605 | Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism]. |
| pfam01204 | Trehalase | 6.52e-17 | 381 | 663 | 222 | 505 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
| COG1626 | TreA | 1.84e-11 | 470 | 663 | 354 | 549 | Neutral trehalase [Carbohydrate transport and metabolism]. |
| PLN02567 | PLN02567 | 1.40e-09 | 357 | 663 | 191 | 541 | alpha,alpha-trehalase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUU00825.1 | 0.0 | 10 | 665 | 16 | 668 |
| QQA30345.1 | 0.0 | 10 | 665 | 16 | 668 |
| QUT61859.1 | 0.0 | 10 | 665 | 16 | 668 |
| BBK86094.1 | 0.0 | 10 | 665 | 16 | 668 |
| ABR38022.1 | 1.94e-299 | 14 | 665 | 18 | 688 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3W7S_A | 3.10e-99 | 35 | 657 | 1 | 747 | Escherichiacoli K12 YgjK complexed with glucose [Escherichia coli K-12],3W7S_B Escherichia coli K12 YgjK complexed with glucose [Escherichia coli K-12],3W7T_A Escherichia coli K12 YgjK complexed with mannose [Escherichia coli K-12],3W7T_B Escherichia coli K12 YgjK complexed with mannose [Escherichia coli K-12],3W7U_A Escherichia coli K12 YgjK complexed with galactose [Escherichia coli K-12],3W7U_B Escherichia coli K12 YgjK complexed with galactose [Escherichia coli K-12] |
| 3W7X_A | 1.66e-98 | 35 | 657 | 1 | 747 | Crystalstructure of E. coli YgjK D324N complexed with melibiose [Escherichia coli K-12],3W7X_B Crystal structure of E. coli YgjK D324N complexed with melibiose [Escherichia coli K-12],5CA3_A Crystal structure of the glycosynthase mutant D324N of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12],5CA3_B Crystal structure of the glycosynthase mutant D324N of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12] |
| 3W7W_A | 2.32e-98 | 35 | 657 | 1 | 747 | Crystalstructure of E. coli YgjK E727A complexed with 2-O-alpha-D-glucopyranosyl-alpha-D-galactopyranose [Escherichia coli K-12],3W7W_B Crystal structure of E. coli YgjK E727A complexed with 2-O-alpha-D-glucopyranosyl-alpha-D-galactopyranose [Escherichia coli K-12],5GW7_A Crystal structure of the glycosynthase mutant E727A of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12],5GW7_B Crystal structure of the glycosynthase mutant E727A of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12] |
| 3D3I_A | 7.67e-94 | 32 | 657 | 5 | 748 | Crystalstructural of Escherichia coli K12 YgjK, a glucosidase belonging to glycoside hydrolase family 63 [Escherichia coli K-12],3D3I_B Crystal structural of Escherichia coli K12 YgjK, a glucosidase belonging to glycoside hydrolase family 63 [Escherichia coli K-12] |
| 7PQQ_B | 5.37e-48 | 436 | 657 | 41 | 296 | ChainB, Anti-RON nanobody,Megabody 91,Glucosidase YgjK [Lama glama] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P42592 | 1.04e-98 | 26 | 657 | 15 | 770 | Glucosidase YgjK OS=Escherichia coli (strain K12) OX=83333 GN=ygjK PE=1 SV=1 |
| D8QTR2 | 4.70e-22 | 326 | 665 | 98 | 487 | Mannosylglycerate hydrolase MGH1 OS=Selaginella moellendorffii OX=88036 GN=MGH PE=1 SV=1 |
| D8T3S4 | 8.37e-22 | 326 | 666 | 98 | 488 | Mannosylglycerate hydrolase MGH2 OS=Selaginella moellendorffii OX=88036 GN=SELMODRAFT_447962 PE=3 SV=1 |
| K5BDL0 | 1.01e-12 | 325 | 663 | 39 | 445 | Glucosylglycerate hydrolase OS=Mycolicibacterium hassiacum (strain DSM 44199 / CIP 105218 / JCM 12690 / 3849) OX=1122247 GN=ggh PE=1 SV=1 |
| Q757L1 | 1.60e-08 | 452 | 644 | 456 | 667 | Probable trehalase OS=Ashbya gossypii (strain ATCC 10895 / CBS 109.51 / FGSC 9923 / NRRL Y-1056) OX=284811 GN=NTH2 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
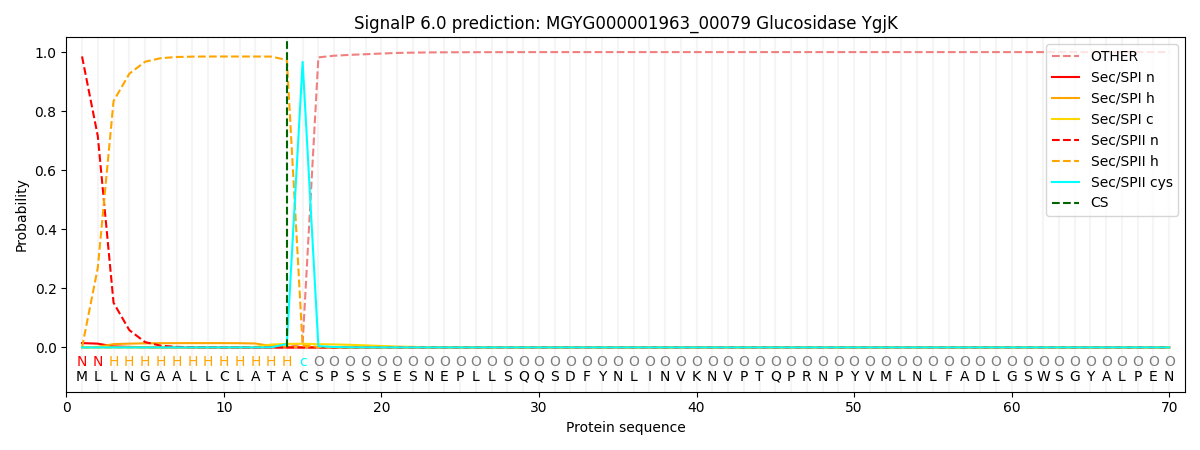
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000046 | 0.014007 | 0.985952 | 0.000009 | 0.000014 | 0.000009 |
