You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001712_03464
You are here: Home > Sequence: MGYG000001712_03464
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacillus_A thuringiensis_S | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales; Bacillaceae_G; Bacillus_A; Bacillus_A thuringiensis_S | |||||||||||
| CAZyme ID | MGYG000001712_03464 | |||||||||||
| CAZy Family | AA10 | |||||||||||
| CAZyme Description | GlcNAc-binding protein A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3413953; End: 3415329 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA10 | 41 | 208 | 4.5e-55 | 0.9943820224719101 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3397 | COG3397 | 1.13e-75 | 15 | 302 | 5 | 300 | Predicted carbohydrate-binding protein, contains CBM5 and CBM33 domains [General function prediction only]. |
| PRK13211 | PRK13211 | 2.99e-75 | 23 | 202 | 7 | 186 | N-acetylglucosamine-binding protein GbpA. |
| cd21177 | LPMO_AA10 | 3.13e-63 | 41 | 202 | 1 | 174 | lytic polysaccharide monooxygenase (LPMO) auxiliary activity family 10 (AA10). AA10 proteins are copper-dependent lytic polysaccharide monooxygenases (LPMOs), which may act on chitin or cellulose. The family used to be called CBM33. Activities in this family include lytic cellulose monooxygenase (C1-hydroxylating) (EC 1.14.99.54), lytic cellulose monooxygenase (C4-dehydrogenating) (EC 1.14.99.56), lytic chitin monooxygenase (EC 1.14.99.53), and lytic xylan monooxygenase/xylan oxidase (glycosidic bond-cleaving) (EC 1.14.99.-). Also included are viral chitin-binding glycoproteins such as fusolin and spheroidin-like proteins. |
| pfam03067 | LPMO_10 | 1.34e-56 | 41 | 202 | 1 | 181 | Lytic polysaccharide mono-oxygenase, cellulose-degrading. This domain is found associated with a wide variety of cellulose binding domains. This is a family of two very closely related proteins that together act as both a C1- and a C4-oxidising lytic polysaccharide mono-oxygenase, degrading cellulose. This domain is also found in baculoviral spheroidins and spindolins, protein of unknown function. |
| COG3401 | FN3 | 4.09e-13 | 219 | 399 | 164 | 340 | Fibronectin type 3 domain [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QBC57830.1 | 0.0 | 1 | 458 | 1 | 458 |
| QRY16100.1 | 0.0 | 1 | 458 | 1 | 458 |
| AND25242.1 | 0.0 | 1 | 458 | 1 | 458 |
| AJH05352.1 | 0.0 | 1 | 458 | 1 | 458 |
| ACK93383.1 | 0.0 | 1 | 458 | 1 | 458 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5LW4_A | 2.11e-94 | 41 | 211 | 1 | 171 | NMRsolution structure of the apo-form of the chitin-active lytic polysaccharide monooxygenase BlLPMO10A [Bacillus licheniformis],6TWE_A Cu(I) NMR solution structure of the chitin-active lytic polysaccharide monooxygenase BlLPMO10A [Bacillus licheniformis DSM 13 = ATCC 14580] |
| 2YOX_A | 4.55e-72 | 41 | 209 | 1 | 177 | Bacillusamyloliquefaciens CBM33 in complex with Cu(I) after photoreduction [Bacillus amyloliquefaciens],2YOX_B Bacillus amyloliquefaciens CBM33 in complex with Cu(I) after photoreduction [Bacillus amyloliquefaciens],2YOY_A Bacillus amyloliquefaciens CBM33 in complex with Cu(I) reduced using ascorbate [Bacillus amyloliquefaciens],2YOY_B Bacillus amyloliquefaciens CBM33 in complex with Cu(I) reduced using ascorbate [Bacillus amyloliquefaciens] |
| 2YOW_A | 4.71e-72 | 41 | 209 | 1 | 177 | Bacillusamyloliquefaciens CBM33 [Bacillus amyloliquefaciens],2YOW_B Bacillus amyloliquefaciens CBM33 [Bacillus amyloliquefaciens],5IJU_A Structure of an AA10 Lytic Polysaccharide Monooxygenase from Bacillus amyloliquefaciens with Cu(II) bound [Bacillus amyloliquefaciens],5IJU_B Structure of an AA10 Lytic Polysaccharide Monooxygenase from Bacillus amyloliquefaciens with Cu(II) bound [Bacillus amyloliquefaciens] |
| 5WSZ_A | 5.78e-67 | 41 | 212 | 1 | 169 | Crystalstructure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki],5WSZ_B Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki],5WSZ_C Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki],5WSZ_D Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis [Bacillus thuringiensis serovar kurstaki] |
| 5L2V_A | 3.41e-61 | 41 | 211 | 1 | 166 | Catalyticdomain of LPMO Lmo2467 from Listeria monocytogenes [Listeria monocytogenes 10403S],5L2V_B Catalytic domain of LPMO Lmo2467 from Listeria monocytogenes [Listeria monocytogenes 10403S] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8EHY2 | 1.19e-50 | 32 | 208 | 19 | 194 | GlcNAc-binding protein A OS=Shewanella oneidensis (strain MR-1) OX=211586 GN=gbpA PE=3 SV=2 |
| Q87FT0 | 2.98e-47 | 30 | 207 | 13 | 199 | GlcNAc-binding protein A OS=Vibrio parahaemolyticus serotype O3:K6 (strain RIMD 2210633) OX=223926 GN=gbpA PE=3 SV=1 |
| Q838S1 | 4.64e-47 | 33 | 209 | 21 | 193 | Lytic chitin monooxygenase OS=Enterococcus faecalis (strain ATCC 700802 / V583) OX=226185 GN=EF_0362 PE=1 SV=1 |
| Q8D7V4 | 1.48e-46 | 30 | 207 | 13 | 198 | GlcNAc-binding protein A OS=Vibrio vulnificus (strain CMCP6) OX=216895 GN=gbpA PE=3 SV=1 |
| Q7MEW9 | 2.06e-46 | 30 | 207 | 13 | 198 | GlcNAc-binding protein A OS=Vibrio vulnificus (strain YJ016) OX=196600 GN=gbpA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
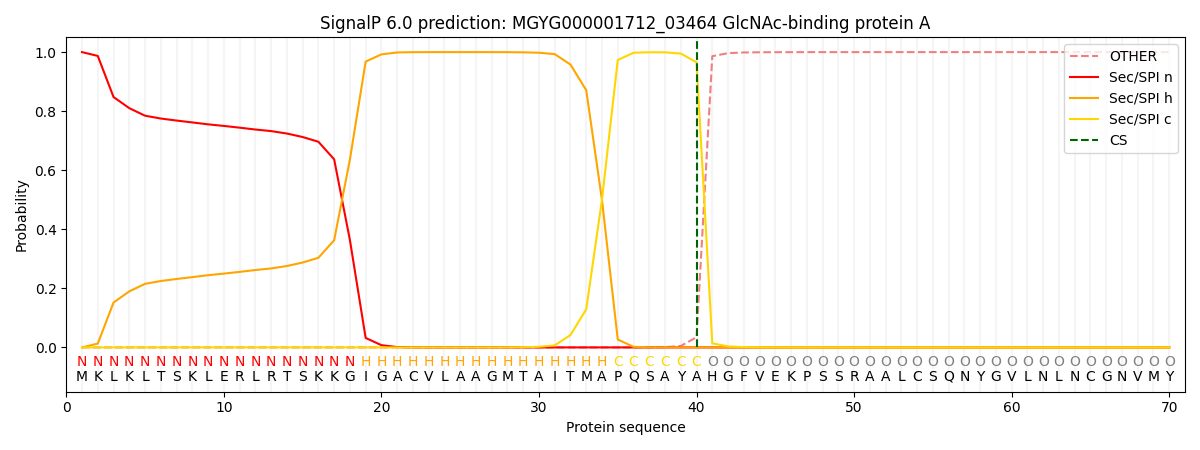
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000400 | 0.998863 | 0.000195 | 0.000198 | 0.000165 | 0.000149 |
