You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001562_00431
You are here: Home > Sequence: MGYG000001562_00431
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes timonensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes timonensis | |||||||||||
| CAZyme ID | MGYG000001562_00431 | |||||||||||
| CAZy Family | GH63 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 280714; End: 282327 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03200 | Glyco_hydro_63 | 9.36e-09 | 332 | 443 | 364 | 487 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| COG3408 | GDB1 | 1.21e-06 | 119 | 459 | 290 | 613 | Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism]. |
| PRK10137 | PRK10137 | 2.48e-05 | 289 | 425 | 614 | 759 | alpha-glucosidase; Provisional |
| COG4354 | COG4354 | 4.89e-05 | 123 | 427 | 381 | 691 | Uncharacterized protein, contains GBA2_N and DUF608 domains [Function unknown]. |
| pfam01204 | Trehalase | 0.002 | 289 | 422 | 341 | 473 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFL78925.1 | 0.0 | 5 | 537 | 5 | 537 |
| BBK99902.1 | 0.0 | 5 | 537 | 5 | 537 |
| BBL10718.1 | 0.0 | 5 | 537 | 5 | 537 |
| BBL07927.1 | 0.0 | 5 | 537 | 5 | 537 |
| BBL07019.1 | 0.0 | 5 | 537 | 3 | 535 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7T66_A | 3.45e-15 | 229 | 443 | 585 | 806 | ChainA, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495],7T66_B Chain B, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495],7T68_A Chain A, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495],7T68_B Chain B, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495],7T6W_A Chain A, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495],7T6W_B Chain B, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495],7T8V_A Chain A, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495],7T8V_B Chain B, Chaetomium alpha glucosidase [Thermochaetoides thermophila DSM 1495] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P94250 | 1.94e-25 | 141 | 478 | 78 | 416 | Uncharacterized protein BB_0381 OS=Borreliella burgdorferi (strain ATCC 35210 / DSM 4680 / CIP 102532 / B31) OX=224326 GN=BB_0381 PE=4 SV=2 |
| O14255 | 6.51e-12 | 193 | 443 | 524 | 799 | Probable mannosyl-oligosaccharide glucosidase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC6G10.09 PE=3 SV=1 |
| D8QTR2 | 9.13e-06 | 350 | 424 | 395 | 463 | Mannosylglycerate hydrolase MGH1 OS=Selaginella moellendorffii OX=88036 GN=MGH PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
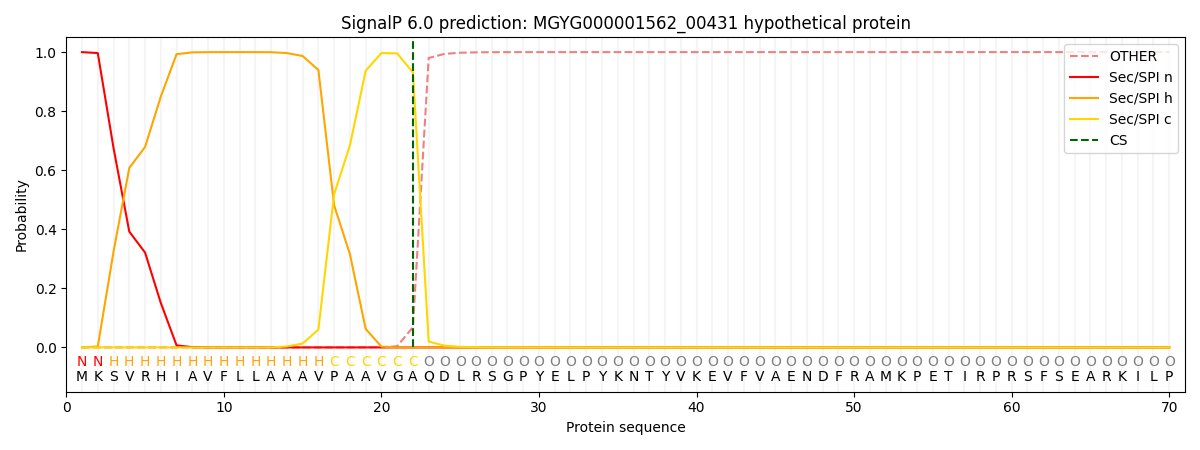
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000325 | 0.998971 | 0.000185 | 0.000167 | 0.000162 | 0.000148 |
