You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001523_01073
You are here: Home > Sequence: MGYG000001523_01073
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
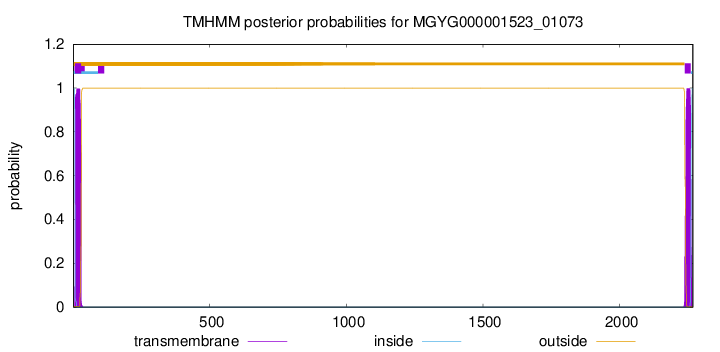
TMHMM annotations
Basic Information help
| Species | Gracilibacillus massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales_D; Amphibacillaceae; Gracilibacillus; Gracilibacillus massiliensis | |||||||||||
| CAZyme ID | MGYG000001523_01073 | |||||||||||
| CAZy Family | GH59 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 299428; End: 306237 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH59 | 52 | 785 | 5.5e-149 | 0.9873217115689382 |
| GH43 | 1664 | 1878 | 1.6e-79 | 0.9904306220095693 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd08983 | GH43_Bt3655-like | 4.15e-115 | 1651 | 1926 | 2 | 260 | Glycosyl hydrolase family 43 protein such as Bacteroides thetaiotaomicron VPI-5482 arabinofuranosidase Bt3655. This glycosyl hydrolase family 43 (GH43)-like family includes the characterized arabinofuranosidases (EC 3.2.1.55): Bacteroides thetaiotaomicron VPI-5482 (Bt3655;BT_3655) and Penicillium chrysogenum 31B Abf43B, as well as Bifidobacterium adolescentis ATCC 15703 beta-xylosidase (EC 3.2.1.37) BAD_1527. It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43 includes enzymes with beta-xylosidase (EC 3.2.1.37), beta-1,3-xylosidase (EC 3.2.1.-), alpha-L-arabinofuranosidase (EC 3.2.1.55), arabinanase (EC 3.2.1.99), xylanase (EC 3.2.1.8), endo-alpha-L-arabinanases (beta-xylanases) and galactan 1,3-beta-galactosidase (EC 3.2.1.145) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| pfam02057 | Glyco_hydro_59 | 5.28e-86 | 54 | 392 | 1 | 290 | Glycosyl hydrolase family 59. |
| cd08978 | GH_F | 4.49e-21 | 1668 | 1924 | 1 | 247 | Glycosyl hydrolase families 43 and 62 form CAZY clan GH-F. This glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) includes family 43 (GH43) and 62 (GH62). GH43 includes enzymes with beta-xylosidase (EC 3.2.1.37), beta-1,3-xylosidase (EC 3.2.1.-), alpha-L-arabinofuranosidase (EC 3.2.1.55), arabinanase (EC 3.2.1.99), xylanase (EC 3.2.1.8), endo-alpha-L-arabinanases (beta-xylanases) and galactan 1,3-beta-galactosidase (EC 3.2.1.145) activities. GH62 includes enzymes characterized as arabinofuranosidases (alpha-L-arabinofuranosidases; EC 3.2.1.55) that specifically cleave either alpha-1,2 or alpha-1,3-L-arabinofuranose side chains from xylans. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many of the enzymes in this family display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. GH62 are also predicted to be inverting enzymes. A common structural feature of both, GH43 and GH62 enzymes, is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| COG5492 | YjdB | 1.68e-13 | 1930 | 2056 | 171 | 295 | Uncharacterized conserved protein YjdB, contains Ig-like domain [General function prediction only]. |
| pfam02368 | Big_2 | 6.05e-13 | 1941 | 2017 | 1 | 77 | Bacterial Ig-like domain (group 2). This family consists of bacterial domains with an Ig-like fold. Members of this family are found in bacterial and phage surface proteins such as intimins. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QGH33896.1 | 0.0 | 4 | 1934 | 5 | 1629 |
| AYA75976.1 | 0.0 | 4 | 1190 | 5 | 1190 |
| AFH63037.1 | 0.0 | 4 | 1193 | 5 | 1195 |
| AFC30718.1 | 0.0 | 4 | 1193 | 5 | 1195 |
| AEI43030.1 | 0.0 | 4 | 1193 | 5 | 1195 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4CCC_A | 1.45e-22 | 46 | 670 | 16 | 539 | StructureOf Mouse Galactocerebrosidase With 4nbdg: Enzyme-substrate Complex [Mus musculus],4CCD_A Structure Of Mouse Galactocerebrosidase With D-galactal: Enzyme-intermediate Complex [Mus musculus],4CCE_A Structure Of Mouse Galactocerebrosidase With Galactose: Enzyme-product Complex [Mus musculus],4UFH_A Mouse Galactocerebrosidase complexed with iso-galacto-fagomine IGF [Mus musculus],4UFI_A Mouse Galactocerebrosidase complexed with aza-galacto-fagomine AGF [Mus musculus],4UFJ_A Mouse Galactocerebrosidase complexed with iso-galacto-fagomine lactam IGL [Mus musculus],4UFK_A Mouse Galactocerebrosidase complexed with dideoxy-imino-lyxitol DIL [Mus musculus],4UFL_A Mouse Galactocerebrosidase complexed with deoxy-galacto-noeurostegine DGN [Mus musculus],4UFM_A Mouse Galactocerebrosidase complexed with 1-deoxy-galacto-nojirimycin DGJ [Mus musculus],5NXB_A Mouse galactocerebrosidase in complex with saposin A [Mus musculus],5NXB_B Mouse galactocerebrosidase in complex with saposin A [Mus musculus],6Y6S_A Chain A, Galactocerebrosidase [Mus musculus],6Y6T_A Chain A, Galactocerebrosidase [Mus musculus] |
| 3ZR5_A | 1.46e-22 | 46 | 670 | 18 | 541 | STRUCTUREOF GALACTOCEREBROSIDASE FROM MOUSE [Mus musculus],3ZR6_A STRUCTURE OF GALACTOCEREBROSIDASE FROM MOUSE IN COMPLEX WITH GALACTOSE [Mus musculus] |
| 5HO0_A | 5.79e-10 | 1523 | 1670 | 552 | 701 | Crystalstructure of AbnA (closed conformation), a GH43 extracellular arabinanase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],5HO2_A Crystal structure of AbnA (open conformation), a GH43 extracellular arabinanase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],5HOF_A Crystal structure of AbnA, a GH43 extracellular arabinanase from Geobacillus stearothermophilus, in complex with arabinopentaose [Geobacillus stearothermophilus],5HP6_A Structure of AbnA, a GH43 extracellular arabinanase from Geobacillus stearothermophilus (a new conformational state) [Geobacillus stearothermophilus] |
| 5HO9_A | 5.69e-09 | 1523 | 1603 | 552 | 632 | Structureof truncated AbnA (domains 1-3), a GH43 arabinanase from Geobacilllus stearothermophilus, in complex with arabinooctaose [Geobacillus stearothermophilus],5HO9_B Structure of truncated AbnA (domains 1-3), a GH43 arabinanase from Geobacilllus stearothermophilus, in complex with arabinooctaose [Geobacillus stearothermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0A3R0A696 | 7.07e-63 | 1245 | 1910 | 83 | 783 | Alpha-L-arabinofuranosidase OS=Bifidobacterium longum subsp. longum (strain ATCC 15707 / DSM 20219 / JCM 1217 / NCTC 11818 / E194b) OX=565042 GN=blArafA PE=1 SV=1 |
| P54803 | 3.18e-23 | 46 | 670 | 46 | 570 | Galactocerebrosidase OS=Homo sapiens OX=9606 GN=GALC PE=1 SV=3 |
| P54804 | 5.22e-23 | 46 | 670 | 30 | 554 | Galactocerebrosidase OS=Canis lupus familiaris OX=9615 GN=GALC PE=1 SV=1 |
| Q0VA39 | 2.85e-22 | 54 | 670 | 47 | 561 | Galactocerebrosidase OS=Xenopus tropicalis OX=8364 GN=galc PE=2 SV=1 |
| O02791 | 3.85e-22 | 46 | 670 | 46 | 570 | Galactocerebrosidase OS=Macaca mulatta OX=9544 GN=GALC PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
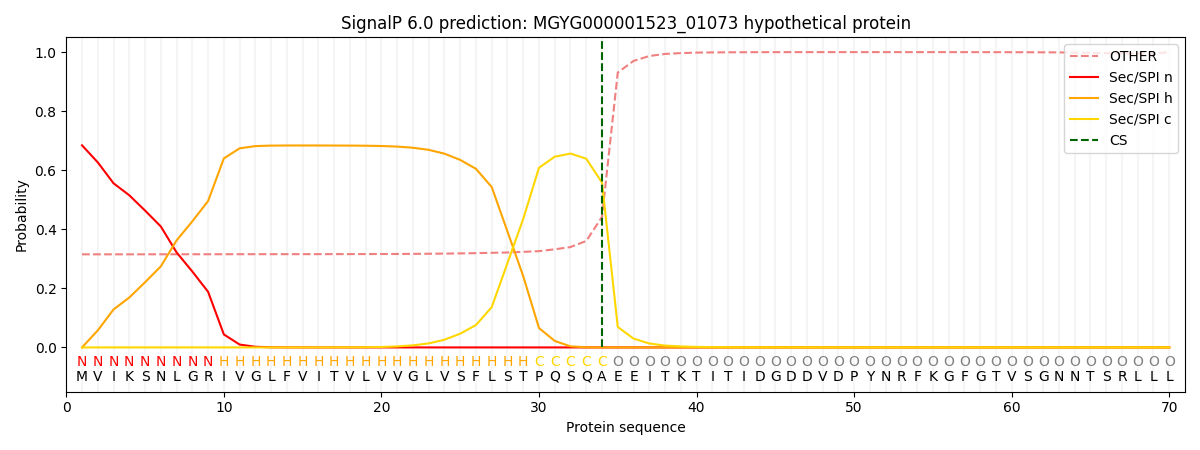
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.337731 | 0.658883 | 0.001194 | 0.000736 | 0.000631 | 0.000815 |