You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001486_01825
You are here: Home > Sequence: MGYG000001486_01825
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Mobilicoccus massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Dermatophilaceae; Mobilicoccus; Mobilicoccus massiliensis | |||||||||||
| CAZyme ID | MGYG000001486_01825 | |||||||||||
| CAZy Family | CE14 | |||||||||||
| CAZyme Description | Mycothiol S-conjugate amidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 279167; End: 281962 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CBM9 | 595 | 787 | 6.1e-36 | 0.989010989010989 |
| CE14 | 62 | 180 | 3.5e-21 | 0.9354838709677419 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd09619 | CBM9_like_4 | 3.02e-40 | 592 | 784 | 1 | 181 | DOMON-like type 9 carbohydrate binding module. Family 9 carbohydrate-binding modules (CBM9) play a role in the microbial degradation of cellulose and hemicellulose (materials found in plants). The domain has previously been called cellulose-binding domain. The polysaccharide binding sites of CBMs with available 3D structure have been found to be either flat surfaces with interactions formed by predominantly aromatic residues (tryptophan and tyrosine), or extended shallow grooves. CBM9 domains found in this uncharacterized heterogeneous subfamily are often located at the C-terminus of longer proteins and may co-occur with various other domains. |
| pfam06452 | CBM9_1 | 1.24e-29 | 595 | 789 | 1 | 182 | Carbohydrate family 9 binding domain-like. CBM9_1 is a C-terminal domain on bacterial xylanase proteins, and it is tandemly repeated in a number of family-members. The CBM9 module binds to amorphous and crystalline cellulose and a range of soluble di- and monosaccharides as well as to cello- and xylo- oligomers of different degrees of polymerization. Comparison of the glucose and cellobiose complexes during crystallisation reveals surprising differences in binding of these two substrates by CBM9-2. Cellobiose was found to bind in a distinct orientation from glucose, while still maintaining optimal stacking and electrostatic interactions with the reducing end sugar. |
| pfam02585 | PIG-L | 8.29e-26 | 64 | 197 | 2 | 125 | GlcNAc-PI de-N-acetylase. Members of this family are related to PIG-L an N-acetylglucosaminylphosphatidylinositol de-N-acetylase (EC:3.5.1.89) that catalyzes the second step in GPI biosynthesis. |
| COG2120 | LmbE | 1.09e-18 | 62 | 212 | 14 | 150 | N-acetylglucosaminyl deacetylase, LmbE family [Carbohydrate transport and metabolism]. |
| cd00005 | CBM9_like_1 | 1.76e-14 | 589 | 787 | 5 | 183 | DOMON-like type 9 carbohydrate binding module of xylanases. Family 9 carbohydrate-binding modules (CBM9) play a role in the microbial degradation of cellulose and hemicellulose (materials found in plants). The domain has previously been called cellulose-binding domain. The polysaccharide binding sites of CBMs with available 3D structure have been found to be either flat surfaces with interactions formed by predominantly aromatic residues (tryptophan and tyrosine), or extended shallow grooves. The CBM9 domain frequently occurs in tandem repeats; members found in this subfamily typically co-occur with glycosyl hydrolase family 10 domains and are annotated as endo-1,4-beta-xylanases. CBM9 from Thermotoga maritima xylanase 10A is reported to have specificity for polysaccharide reducing ends. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QLD26882.1 | 6.77e-261 | 53 | 930 | 39 | 905 |
| QDY10008.1 | 6.25e-254 | 38 | 929 | 17 | 896 |
| BCL13207.1 | 9.13e-243 | 44 | 831 | 24 | 809 |
| AYY14116.1 | 1.24e-241 | 58 | 930 | 65 | 937 |
| QZZ32409.1 | 1.41e-116 | 49 | 830 | 18 | 559 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D6Y7M2 | 2.64e-09 | 58 | 229 | 3 | 180 | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase OS=Thermobispora bispora (strain ATCC 19993 / DSM 43833 / CBS 139.67 / JCM 10125 / KCTC 9307 / NBRC 14880 / R51) OX=469371 GN=mshB PE=3 SV=1 |
| Q82IJ5 | 4.38e-09 | 58 | 227 | 6 | 183 | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase OS=Streptomyces avermitilis (strain ATCC 31267 / DSM 46492 / JCM 5070 / NBRC 14893 / NCIMB 12804 / NRRL 8165 / MA-4680) OX=227882 GN=mshB PE=3 SV=1 |
| D2S9Y0 | 1.05e-08 | 62 | 283 | 8 | 247 | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase OS=Geodermatophilus obscurus (strain ATCC 25078 / DSM 43160 / JCM 3152 / KCC A-0152 / KCTC 9177 / NBRC 13315 / NRRL B-3577 / G-20) OX=526225 GN=mshB PE=3 SV=1 |
| C9ZBY0 | 3.11e-07 | 62 | 227 | 23 | 196 | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase OS=Streptomyces scabiei (strain 87.22) OX=680198 GN=mshB PE=3 SV=1 |
| C1AZH2 | 1.42e-06 | 62 | 241 | 7 | 193 | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase OS=Rhodococcus opacus (strain B4) OX=632772 GN=mshB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as TAT
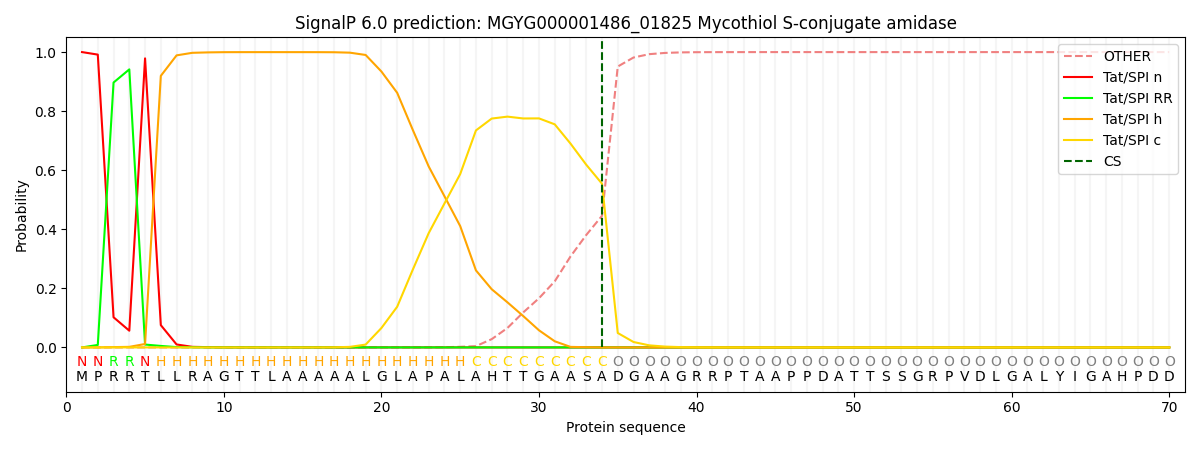
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 0.000000 | 1.000017 | 0.000000 | 0.000000 |
