You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001422_01305
You are here: Home > Sequence: MGYG000001422_01305
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides oleiciplenus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides oleiciplenus | |||||||||||
| CAZyme ID | MGYG000001422_01305 | |||||||||||
| CAZy Family | GH29 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1428866; End: 1432993 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH29 | 20 | 344 | 3.2e-92 | 0.9017341040462428 |
| PL8 | 952 | 1214 | 1.5e-47 | 0.9959514170040485 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00812 | Alpha_L_fucos | 3.30e-93 | 28 | 384 | 20 | 384 | Alpha-L-fucosidase. O-Glycosyl hydrolases (EC 3.2.1.-) are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycosyl hydrolases, based on sequence similarity, has led to the definition of 85 different families. This classification is available on the CAZy (CArbohydrate-Active EnZymes) web site. Because the fold of proteins is better conserved than their sequences, some of the families can be grouped in 'clans'. Family 29 encompasses alpha-L-fucosidases, which is a lysosomal enzyme responsible for hydrolyzing the alpha-1,6-linked fucose joined to the reducing-end N-acetylglucosamine of the carbohydrate moieties of glycoproteins. Deficiency of alpha-L-fucosidase results in the lysosomal storage disease fucosidosis. |
| pfam01120 | Alpha_L_fucos | 2.99e-79 | 28 | 340 | 19 | 333 | Alpha-L-fucosidase. |
| cd01083 | GAG_Lyase | 1.84e-42 | 789 | 1299 | 194 | 693 | Glycosaminoglycan (GAG) polysaccharide lyase family. This family consists of a group of secreted bacterial lyase enzymes capable of acting on glycosaminoglycans, such as hyaluronan and chondroitin, in the extracellular matrix of host tissues, contributing to the invasive capacity of the pathogen. These are broad-specificity glycosaminoglycan lyases which recognize uronyl residues in polysaccharides and cleave their glycosidic bonds via a beta-elimination reaction to form a double bond between C-4 and C-5 of the non-reducing terminal uronyl residues of released products. Substrates include chondroitin, chondroitin 4-sulfate, chondroitin 6-sulfate, and hyaluronic acid. Family members include chondroitin AC lyase, chondroitin abc lyase, xanthan lyase, and hyalurate lyase. |
| COG3669 | AfuC | 1.61e-36 | 41 | 344 | 1 | 321 | Alpha-L-fucosidase [Carbohydrate transport and metabolism]. |
| pfam09093 | Lyase_catalyt | 2.20e-20 | 621 | 922 | 32 | 351 | Lyase, catalytic. Members of this family are predominantly found in chondroitin ABC lyase I, and adopt a helical structure, with fifteen alpha-helices which are at least two turns long and several short helical turns. The bulk of the domain is formed by ten alpha-helices forming five hairpin-like pairs and arranged into an incomplete toroid, the (alpha/alpha)5 fold. Additionally, two long and two short alpha-helices at the N-terminus of the domain wrap around the toroid. At the C-terminal end of the toroid there is one additional short alpha-helix. This domain is required for degradation of polysaccharides containing 1,4-beta-D-hexosaminyl and 1,3-beta-D-glucoronosyl or 1,3-alpha-L-iduronosyl linkages to disaccharides containing 4-deoxy-beta-D-gluc-4-enuronosyl groups. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT74184.1 | 1.39e-310 | 10 | 597 | 9 | 609 |
| BCI63816.1 | 3.80e-190 | 591 | 1375 | 203 | 984 |
| BCI64222.1 | 6.42e-171 | 598 | 1363 | 198 | 976 |
| QQE11582.1 | 3.62e-145 | 12 | 594 | 1 | 594 |
| QHI69131.1 | 5.70e-127 | 23 | 586 | 2 | 567 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6GN6_A | 6.96e-74 | 23 | 413 | 26 | 422 | Alpha-L-fucosidaseisoenzyme 1 from Paenibacillus thiaminolyticus [Paenibacillus thiaminolyticus],6GN6_B Alpha-L-fucosidase isoenzyme 1 from Paenibacillus thiaminolyticus [Paenibacillus thiaminolyticus],6GN6_C Alpha-L-fucosidase isoenzyme 1 from Paenibacillus thiaminolyticus [Paenibacillus thiaminolyticus],6GN6_D Alpha-L-fucosidase isoenzyme 1 from Paenibacillus thiaminolyticus [Paenibacillus thiaminolyticus],6GN6_E Alpha-L-fucosidase isoenzyme 1 from Paenibacillus thiaminolyticus [Paenibacillus thiaminolyticus],6GN6_F Alpha-L-fucosidase isoenzyme 1 from Paenibacillus thiaminolyticus [Paenibacillus thiaminolyticus] |
| 7DB5_A | 1.95e-67 | 31 | 561 | 37 | 578 | ChainA, Alpha-L-fucosidase [Vibrio sp. EJY3] |
| 6O18_A | 1.06e-62 | 30 | 321 | 7 | 325 | ChainA, AlfC [Lacticaseibacillus casei],6O18_B Chain B, AlfC [Lacticaseibacillus casei],6O18_C Chain C, AlfC [Lacticaseibacillus casei],6O18_D Chain D, AlfC [Lacticaseibacillus casei],6O1A_A Chain A, AlfC [Lacticaseibacillus casei],6O1A_B Chain B, AlfC [Lacticaseibacillus casei],6O1A_C Chain C, AlfC [Lacticaseibacillus casei],6O1A_D Chain D, AlfC [Lacticaseibacillus casei] |
| 6O1I_A | 1.44e-62 | 30 | 321 | 7 | 325 | ChainA, AlfC [Lacticaseibacillus casei],6O1I_B Chain B, AlfC [Lacticaseibacillus casei],6O1I_C Chain C, AlfC [Lacticaseibacillus casei],6O1I_D Chain D, AlfC [Lacticaseibacillus casei] |
| 6O1C_A | 1.24e-61 | 30 | 321 | 7 | 325 | ChainA, AlfC [Lacticaseibacillus casei],6O1C_B Chain B, AlfC [Lacticaseibacillus casei],6O1C_C Chain C, AlfC [Lacticaseibacillus casei],6O1C_D Chain D, AlfC [Lacticaseibacillus casei],6O1C_E Chain E, AlfC [Lacticaseibacillus casei],6O1C_F Chain F, AlfC [Lacticaseibacillus casei],6O1C_G Chain G, AlfC [Lacticaseibacillus casei],6O1C_H Chain H, AlfC [Lacticaseibacillus casei],6OHE_A Chain A, AlfC [Lacticaseibacillus casei],6OHE_B Chain B, AlfC [Lacticaseibacillus casei],6OHE_C Chain C, AlfC [Lacticaseibacillus casei],6OHE_D Chain D, AlfC [Lacticaseibacillus casei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P59807 | 6.56e-58 | 721 | 1330 | 398 | 987 | Chondroitin sulfate ABC endolyase OS=Proteus vulgaris OX=585 PE=1 SV=2 |
| Q60HF8 | 3.62e-32 | 29 | 382 | 53 | 413 | Tissue alpha-L-fucosidase OS=Macaca fascicularis OX=9541 GN=FUCA1 PE=2 SV=1 |
| P04066 | 8.53e-32 | 30 | 382 | 52 | 411 | Tissue alpha-L-fucosidase OS=Homo sapiens OX=9606 GN=FUCA1 PE=1 SV=4 |
| Q99LJ1 | 1.24e-31 | 29 | 382 | 37 | 397 | Tissue alpha-L-fucosidase OS=Mus musculus OX=10090 GN=Fuca1 PE=1 SV=1 |
| P17164 | 6.36e-31 | 30 | 382 | 48 | 407 | Tissue alpha-L-fucosidase OS=Rattus norvegicus OX=10116 GN=Fuca1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
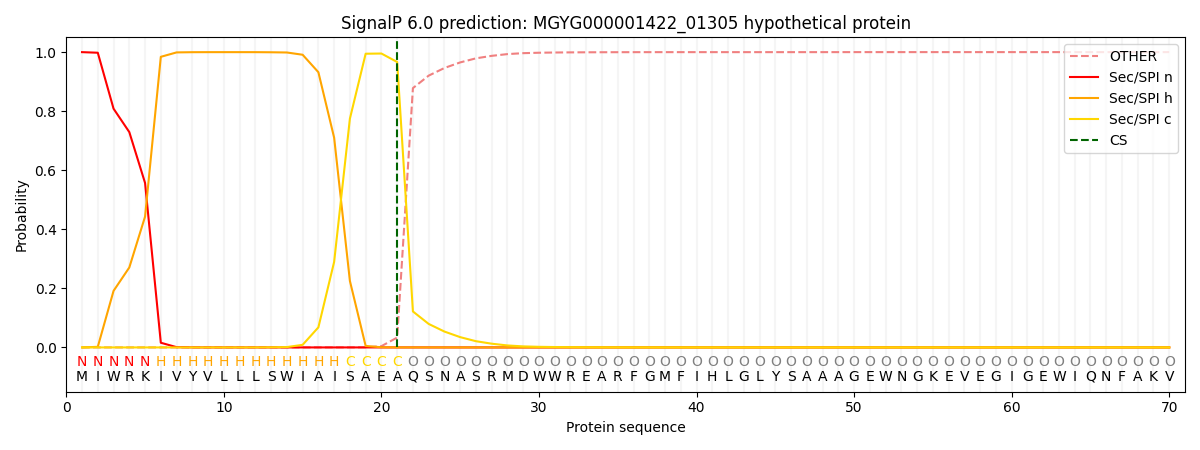
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001184 | 0.997712 | 0.000350 | 0.000266 | 0.000222 | 0.000210 |
