You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001317_02304
You are here: Home > Sequence: MGYG000001317_02304
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Citrobacter portucalensis_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Citrobacter; Citrobacter portucalensis_A | |||||||||||
| CAZyme ID | MGYG000001317_02304 | |||||||||||
| CAZy Family | GH27 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 52877; End: 55795 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH27 | 424 | 666 | 8.9e-16 | 0.925764192139738 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd14791 | GH36 | 8.41e-27 | 298 | 596 | 3 | 296 | glycosyl hydrolase family 36 (GH36). GH36 enzymes occur in prokaryotes, eukaryotes, and archaea with a wide range of hydrolytic activities, including alpha-galactosidase, alpha-N-acetylgalactosaminidase, stachyose synthase, and raffinose synthase. All GH36 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH36 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| cd14792 | GH27 | 7.64e-18 | 298 | 539 | 2 | 215 | glycosyl hydrolase family 27 (GH27). GH27 enzymes occur in eukaryotes, prokaryotes, and archaea with a wide range of hydrolytic activities, including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-N-acetylgalactosaminidase, and 3-alpha-isomalto-dextranase. All GH27 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH27 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| pfam16499 | Melibiase_2 | 2.13e-05 | 318 | 440 | 33 | 132 | Alpha galactosidase A. |
| PLN02692 | PLN02692 | 1.93e-04 | 298 | 481 | 57 | 205 | alpha-galactosidase |
| PLN02808 | PLN02808 | 0.001 | 327 | 367 | 63 | 116 | alpha-galactosidase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AYN29307.1 | 0.0 | 19 | 969 | 18 | 967 |
| AJQ98020.1 | 0.0 | 8 | 969 | 13 | 981 |
| QQE42004.1 | 0.0 | 8 | 969 | 13 | 981 |
| AYQ71140.1 | 5.72e-305 | 23 | 971 | 38 | 1027 |
| ANS75585.1 | 2.57e-256 | 25 | 968 | 40 | 1027 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6LCJ_A | 9.34e-07 | 326 | 519 | 184 | 370 | TtGalA,alpha-galactosidase from Thermus thermopilus in apo form [Thermus thermophilus HB8],6LCJ_B TtGalA, alpha-galactosidase from Thermus thermopilus in apo form [Thermus thermophilus HB8],6LCJ_C TtGalA, alpha-galactosidase from Thermus thermopilus in apo form [Thermus thermophilus HB8],6LCJ_D TtGalA, alpha-galactosidase from Thermus thermopilus in apo form [Thermus thermophilus HB8],6LCJ_E TtGalA, alpha-galactosidase from Thermus thermopilus in apo form [Thermus thermophilus HB8],6LCJ_F TtGalA, alpha-galactosidase from Thermus thermopilus in apo form [Thermus thermophilus HB8],6LCK_A TtGalA, alpha-galactosidase from Thermus thermophilus in complex with p-nitrophenyl alpha-D-galactopyranoside (alpha-NPG) [Thermus thermophilus HB8],6LCK_B TtGalA, alpha-galactosidase from Thermus thermophilus in complex with p-nitrophenyl alpha-D-galactopyranoside (alpha-NPG) [Thermus thermophilus HB8],6LCK_C TtGalA, alpha-galactosidase from Thermus thermophilus in complex with p-nitrophenyl alpha-D-galactopyranoside (alpha-NPG) [Thermus thermophilus HB8],6LCL_A TtGalA, alpha-galactosidase from Thermus thermophilus in complex with stachyose [Thermus thermophilus HB8],6LCL_C TtGalA, alpha-galactosidase from Thermus thermophilus in complex with stachyose [Thermus thermophilus HB8],6LCL_E TtGalA, alpha-galactosidase from Thermus thermophilus in complex with stachyose [Thermus thermophilus HB8] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
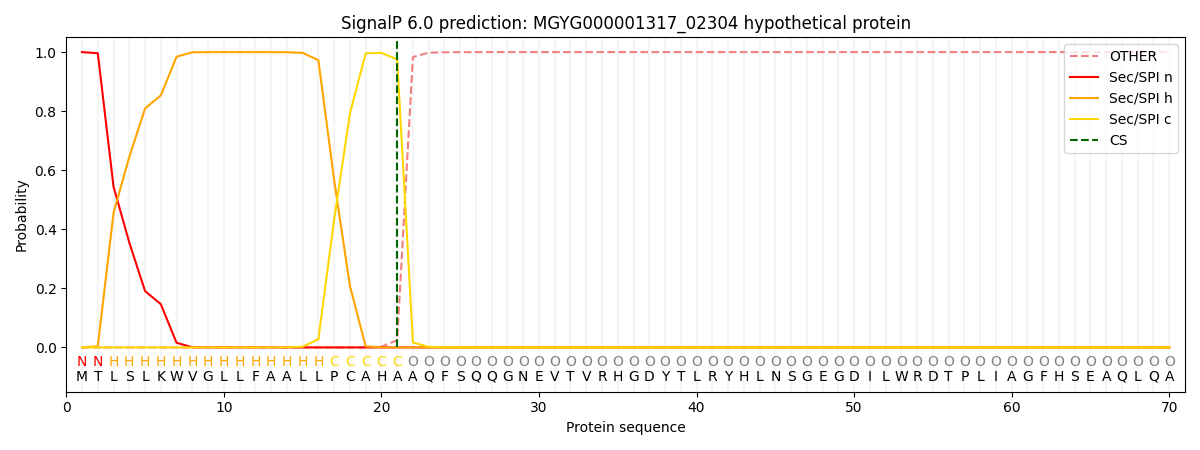
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000301 | 0.998933 | 0.000227 | 0.000188 | 0.000171 | 0.000155 |
