You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001262_01505
You are here: Home > Sequence: MGYG000001262_01505
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Sphingomonas ginsenosidimutans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Alphaproteobacteria; Sphingomonadales; Sphingomonadaceae; Sphingomonas; Sphingomonas ginsenosidimutans | |||||||||||
| CAZyme ID | MGYG000001262_01505 | |||||||||||
| CAZy Family | GH36 | |||||||||||
| CAZyme Description | Alpha-galactosidase AgaA | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 13325; End: 15526 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH36 | 25 | 708 | 1.5e-230 | 0.9941860465116279 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02065 | Melibiase | 2.78e-143 | 278 | 625 | 2 | 347 | Melibiase. Glycoside hydrolase families GH27, GH31 and GH36 form the glycoside hydrolase clan GH-D. Glycoside hydrolase family 36 can be split into 11 families, GH36A to GH36K. This family includes enzymes from GH36A-B and GH36D-K and from GH27. |
| COG3345 | GalA | 1.94e-141 | 24 | 726 | 2 | 686 | Alpha-galactosidase [Carbohydrate transport and metabolism]. |
| cd14791 | GH36 | 1.41e-132 | 319 | 617 | 2 | 299 | glycosyl hydrolase family 36 (GH36). GH36 enzymes occur in prokaryotes, eukaryotes, and archaea with a wide range of hydrolytic activities, including alpha-galactosidase, alpha-N-acetylgalactosaminidase, stachyose synthase, and raffinose synthase. All GH36 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH36 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| pfam16875 | Glyco_hydro_36N | 2.06e-71 | 43 | 272 | 1 | 255 | Glycosyl hydrolase family 36 N-terminal domain. This domain is found at the N-terminus of many family 36 glycoside hydrolases. It has a beta-supersandwich fold. |
| pfam16874 | Glyco_hydro_36C | 6.78e-11 | 643 | 724 | 5 | 78 | Glycosyl hydrolase family 36 C-terminal domain. This domain is found at the C-terminus of many family 36 glycoside hydrolases. It has a beta-sandwich structure with a Greek key motif. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QWC57685.1 | 0.0 | 13 | 726 | 21 | 734 |
| AFE82291.1 | 0.0 | 13 | 723 | 21 | 734 |
| AGH49656.1 | 0.0 | 5 | 723 | 7 | 725 |
| QWF14853.1 | 0.0 | 13 | 723 | 24 | 734 |
| ATE72151.1 | 0.0 | 10 | 723 | 21 | 734 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4FNR_A | 6.70e-140 | 18 | 684 | 4 | 693 | Crystalstructure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNR_B Crystal structure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNR_C Crystal structure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNR_D Crystal structure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
| 4FNQ_A | 3.71e-139 | 18 | 684 | 4 | 693 | Crystalstructure of GH36 alpha-galactosidase AgaB from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
| 4FNP_A | 7.35e-139 | 18 | 684 | 4 | 693 | Crystalstructure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNP_B Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNP_C Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNP_D Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNS_A Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus],4FNS_B Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus],4FNS_C Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus],4FNS_D Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus] |
| 4FNT_A | 2.05e-138 | 18 | 684 | 4 | 693 | Crystalstructure of GH36 alpha-galactosidase AgaA A355E D548N from Geobacillus stearothermophilus in complex with raffinose [Geobacillus stearothermophilus],4FNT_B Crystal structure of GH36 alpha-galactosidase AgaA A355E D548N from Geobacillus stearothermophilus in complex with raffinose [Geobacillus stearothermophilus],4FNT_C Crystal structure of GH36 alpha-galactosidase AgaA A355E D548N from Geobacillus stearothermophilus in complex with raffinose [Geobacillus stearothermophilus],4FNT_D Crystal structure of GH36 alpha-galactosidase AgaA A355E D548N from Geobacillus stearothermophilus in complex with raffinose [Geobacillus stearothermophilus] |
| 4FNU_A | 5.73e-138 | 18 | 684 | 4 | 693 | Crystalstructure of GH36 alpha-galactosidase AgaA A355E D478A from Geobacillus stearothermophilus in complex with stachyose [Geobacillus stearothermophilus],4FNU_B Crystal structure of GH36 alpha-galactosidase AgaA A355E D478A from Geobacillus stearothermophilus in complex with stachyose [Geobacillus stearothermophilus],4FNU_C Crystal structure of GH36 alpha-galactosidase AgaA A355E D478A from Geobacillus stearothermophilus in complex with stachyose [Geobacillus stearothermophilus],4FNU_D Crystal structure of GH36 alpha-galactosidase AgaA A355E D478A from Geobacillus stearothermophilus in complex with stachyose [Geobacillus stearothermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9ALJ4 | 3.67e-139 | 18 | 684 | 4 | 693 | Alpha-galactosidase AgaA OS=Geobacillus stearothermophilus OX=1422 GN=agaA PE=1 SV=1 |
| Q5AU92 | 4.33e-124 | 26 | 726 | 44 | 749 | Alpha-galactosidase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=aglC PE=1 SV=1 |
| G1UB44 | 3.13e-122 | 19 | 687 | 8 | 700 | Alpha-galactosidase Mel36A OS=Lactobacillus acidophilus (strain ATCC 700396 / NCK56 / N2 / NCFM) OX=272621 GN=melA PE=1 SV=1 |
| Q0CEF5 | 5.54e-121 | 20 | 717 | 14 | 715 | Probable alpha-galactosidase G OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=aglG PE=3 SV=1 |
| Q9UUZ4 | 1.50e-119 | 26 | 684 | 41 | 714 | Alpha-galactosidase C OS=Aspergillus niger OX=5061 GN=aglC PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
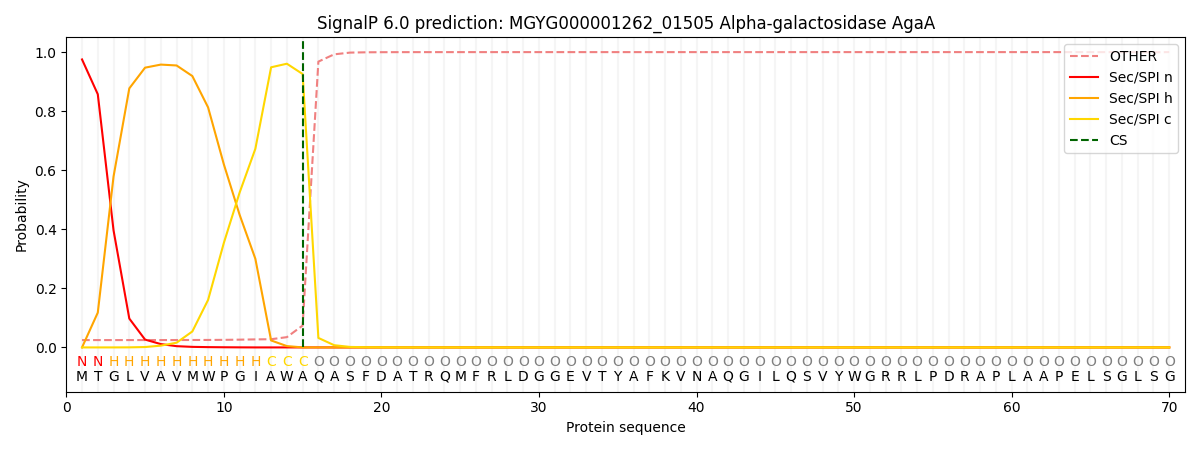
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.028331 | 0.970484 | 0.000291 | 0.000332 | 0.000267 | 0.000250 |
