You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001214_00191
You are here: Home > Sequence: MGYG000001214_00191
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Lancefieldella rimae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Coriobacteriia; Coriobacteriales; Atopobiaceae; Lancefieldella; Lancefieldella rimae | |||||||||||
| CAZyme ID | MGYG000001214_00191 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 21809; End: 22846 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 54 | 323 | 2.9e-39 | 0.9602649006622517 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2730 | BglC | 6.30e-12 | 2 | 234 | 13 | 247 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| pfam00150 | Cellulase | 1.86e-05 | 62 | 282 | 28 | 239 | Cellulase (glycosyl hydrolase family 5). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACV51516.1 | 3.75e-184 | 3 | 345 | 2 | 346 |
| QOY61077.1 | 5.74e-146 | 7 | 344 | 6 | 346 |
| ADK68544.1 | 1.65e-128 | 3 | 345 | 2 | 347 |
| SDR87855.1 | 3.59e-120 | 5 | 344 | 4 | 346 |
| AKT48931.1 | 1.61e-118 | 3 | 345 | 2 | 346 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6ZB9_A | 9.91e-32 | 7 | 331 | 3 | 373 | ChainA, Exo-beta-1,3-glucanase [uncultured bacterium],6ZB9_B Chain B, Exo-beta-1,3-glucanase [uncultured bacterium] |
| 6ZB8_A | 1.90e-31 | 7 | 331 | 3 | 373 | ChainA, Exo-beta-1,3-glucanase variant E167Q/E295Q [uncultured bacterium],6ZB8_B Chain B, Exo-beta-1,3-glucanase variant E167Q/E295Q [uncultured bacterium] |
| 3O6A_A | 1.88e-19 | 5 | 323 | 12 | 366 | F144Y/F258YDouble Mutant of Exo-beta-1,3-glucanase from Candida albicans at 2 A [Candida albicans] |
| 1EQP_A | 2.46e-19 | 5 | 323 | 7 | 361 | Exo-b-(1,3)-glucanaseFrom Candida Albicans [Candida albicans] |
| 3N9K_A | 2.56e-19 | 5 | 323 | 12 | 366 | F229A/E292SDouble Mutant of Exo-beta-1,3-glucanase from Candida albicans in Complex with Laminaritriose at 1.7 A [Candida albicans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A1CRV0 | 8.54e-37 | 4 | 338 | 39 | 398 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=exgA PE=3 SV=2 |
| B0XN12 | 2.33e-36 | 6 | 338 | 41 | 399 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=exgA PE=3 SV=1 |
| Q4WK60 | 1.68e-35 | 6 | 338 | 41 | 399 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=exgA PE=3 SV=1 |
| A1D4Q5 | 6.23e-35 | 6 | 338 | 41 | 399 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=exgA PE=3 SV=1 |
| A2RAR6 | 1.61e-32 | 4 | 338 | 39 | 399 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=exgA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
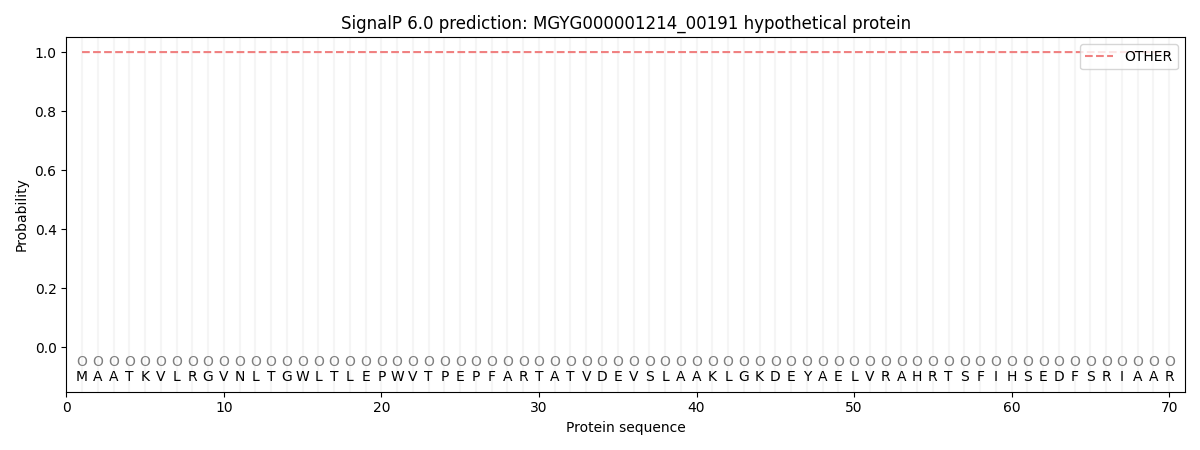
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000064 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
