You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001192_01518
You are here: Home > Sequence: MGYG000001192_01518
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA1777 sp900549865 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; UBA1777; UBA1777 sp900549865 | |||||||||||
| CAZyme ID | MGYG000001192_01518 | |||||||||||
| CAZy Family | GH31 | |||||||||||
| CAZyme Description | Sulfoquinovosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7303; End: 9309 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH31 | 219 | 651 | 2.1e-92 | 0.9742388758782201 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10426 | PRK10426 | 0.0 | 1 | 663 | 6 | 630 | alpha-glucosidase; Provisional |
| cd06594 | GH31_glucosidase_YihQ | 1.58e-176 | 235 | 543 | 1 | 318 | alpha-glucosidase YihQ-like. YihQ is a bacterial alpha-glucosidase with a conserved glycosyl hydrolase family 31 (GH31) domain that catalyzes the release of an alpha-glucosyl residue from the non-reducing end of alpha-glucoside substrates such as alpha-glucosyl fluoride. Orthologs of YihQ that have not yet been functionally characterized are present in plants and fungi. YihQ has sequence similarity to other GH31 enzymes such as CtsZ, a 6-alpha-glucosyltransferase from Bacillus globisporus, and YicI, an alpha-xylosidase from Echerichia coli. These latter two belong to different GH31 subfamilies than YihQ. In bacteria, YihQ (along with YihO) is important for bacterial O-antigen capsule assembly and translocation. |
| COG1501 | YicI | 8.86e-138 | 86 | 651 | 103 | 664 | Alpha-glucosidase, glycosyl hydrolase family GH31 [Carbohydrate transport and metabolism]. |
| pfam01055 | Glyco_hydro_31 | 1.30e-76 | 221 | 650 | 6 | 435 | Glycosyl hydrolases family 31. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. Family 31 comprises of enzymes that are, or similar to, alpha- galactosidases. |
| cd06592 | GH31_NET37 | 9.52e-54 | 241 | 624 | 3 | 364 | glucosidase NET37. NET37 (also known as KIAA1161) is a human lamina-associated nuclear envelope transmembrane protein. A member of the glycosyl hydrolase family 31 (GH31) , it has been shown to be required for myogenic differentiation of C2C12 cells. Related proteins are found in eukaryotes and prokaryotes. Enzymes of the GH31 family possess a wide range of different hydrolytic activities including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase and alpha-1,4-glucan lyase. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALX96444.1 | 5.90e-244 | 4 | 663 | 5 | 672 |
| AHG19153.1 | 1.80e-243 | 4 | 666 | 5 | 675 |
| VEI74313.1 | 2.38e-243 | 4 | 663 | 5 | 672 |
| QXN60498.1 | 2.38e-243 | 4 | 663 | 5 | 672 |
| QKJ58027.1 | 2.38e-243 | 4 | 663 | 5 | 672 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5AED_A | 2.15e-230 | 24 | 665 | 29 | 677 | Abacterial protein structure in glycoside hydrolase family 31 [Escherichia coli K-12],5AED_B A bacterial protein structure in glycoside hydrolase family 31 [Escherichia coli K-12],5AEG_A A bacterial protein structure in glycoside hydrolase family 31. [Escherichia coli K-12],5AEG_B A bacterial protein structure in glycoside hydrolase family 31. [Escherichia coli K-12],5OHT_A A GH31 family sulfoquinovosidase from E. coli in complex with aza-sugar inhibitor IFGSQ [Escherichia coli K-12],5OHT_B A GH31 family sulfoquinovosidase from E. coli in complex with aza-sugar inhibitor IFGSQ [Escherichia coli K-12] |
| 5AEE_A | 1.23e-229 | 24 | 665 | 29 | 677 | Abacterial protein structure in glycoside hydrolase family 31 [Escherichia coli K-12],5AEE_B A bacterial protein structure in glycoside hydrolase family 31 [Escherichia coli K-12] |
| 6PNR_A | 1.15e-227 | 16 | 663 | 11 | 661 | ChainA, Alpha-glucosidase [[Eubacterium] rectale],6PNR_C Chain C, Alpha-glucosidase [[Eubacterium] rectale] |
| 5OHY_A | 9.77e-210 | 17 | 663 | 12 | 660 | ChainA, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHY_B Chain B, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHY_C Chain C, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHY_D Chain D, Alpha-glucosidase yihQ [Agrobacterium tumefaciens] |
| 5OHS_A | 5.56e-209 | 17 | 663 | 12 | 660 | ChainA, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHS_B Chain B, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHS_C Chain C, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHS_D Chain D, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHS_E Chain E, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHS_F Chain F, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHS_G Chain G, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],5OHS_H Chain H, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],7OFX_A Chain A, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],7OFX_B Chain B, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],7OFX_C Chain C, Alpha-glucosidase yihQ [Agrobacterium tumefaciens],7OFX_D Chain D, Alpha-glucosidase yihQ [Agrobacterium tumefaciens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P32138 | 8.98e-230 | 24 | 665 | 29 | 677 | Sulfoquinovosidase OS=Escherichia coli (strain K12) OX=83333 GN=yihQ PE=1 SV=3 |
| B3PEE6 | 4.64e-47 | 69 | 664 | 105 | 679 | Oligosaccharide 4-alpha-D-glucosyltransferase OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=agd31B PE=1 SV=1 |
| P31434 | 1.43e-41 | 106 | 635 | 148 | 646 | Alpha-xylosidase OS=Escherichia coli (strain K12) OX=83333 GN=yicI PE=1 SV=2 |
| D2PPM7 | 4.83e-36 | 56 | 659 | 95 | 718 | 1,3-alpha-isomaltosidase OS=Kribbella flavida (strain DSM 17836 / JCM 10339 / NBRC 14399) OX=479435 GN=Kfla_1895 PE=1 SV=1 |
| Q9F234 | 2.02e-35 | 115 | 650 | 144 | 667 | Alpha-glucosidase 2 OS=Bacillus thermoamyloliquefaciens OX=1425 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
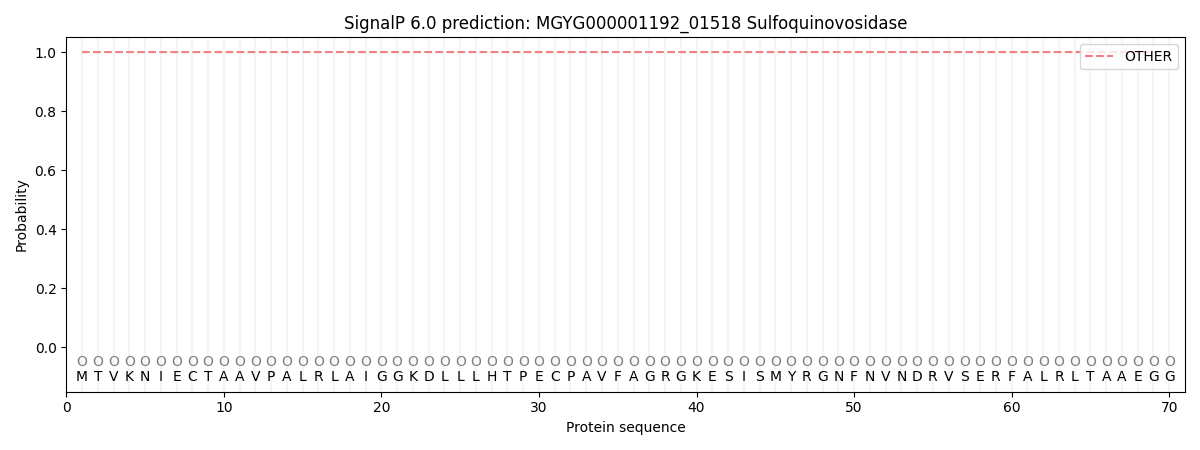
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000065 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
