You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000491_01345
You are here: Home > Sequence: MGYG000000491_01345
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Zag111 sp003258735 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Cyanobacteria; Vampirovibrionia; Gastranaerophilales; Gastranaerophilaceae; Zag111; Zag111 sp003258735 | |||||||||||
| CAZyme ID | MGYG000000491_01345 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | Beta-monoglucosyldiacylglycerol synthase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 206408; End: 207775 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 67 | 291 | 6.4e-39 | 0.9869565217391304 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1215 | BcsA | 7.10e-55 | 18 | 426 | 9 | 424 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| cd06423 | CESA_like | 4.51e-54 | 71 | 250 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| PRK11204 | PRK11204 | 3.06e-42 | 35 | 417 | 20 | 405 | N-glycosyltransferase; Provisional |
| cd06437 | CESA_CaSu_A2 | 1.41e-36 | 67 | 293 | 1 | 231 | Cellulose synthase catalytic subunit A2 (CESA2) is a catalytic subunit or a catalytic subunit substitute of the cellulose synthase complex. Cellulose synthase (CESA) catalyzes the polymerization reaction of cellulose using UDP-glucose as the substrate. Cellulose is an aggregate of unbranched polymers of beta-1,4-linked glucose residues, which is an abundant polysaccharide produced by plants and in varying degrees by several other organisms including algae, bacteria, fungi, and even some animals. Genomes from higher plants harbor multiple CESA genes. There are ten in Arabidopsis. At least three different CESA proteins are required to form a functional complex. In Arabidopsis, CESA1, 3 and 6 and CESA4, 7 and 8, are required for cellulose biosynthesis during primary and secondary cell wall formation. CESA2 is very closely related to CESA6 and is viewed as a prime substitute for CESA6. They functionally compensate each other. The cesa2 and cesa6 double mutant plants were significantly smaller, while the single mutant plants were almost normal. |
| cd06439 | CESA_like_1 | 9.87e-34 | 56 | 275 | 18 | 229 | CESA_like_1 is a member of the cellulose synthase (CESA) superfamily. This is a subfamily of cellulose synthase (CESA) superfamily. CESA superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members of the superfamily include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOR38294.1 | 6.37e-204 | 44 | 434 | 2 | 391 |
| BDA68096.1 | 3.02e-76 | 23 | 426 | 72 | 463 |
| BAZ42219.1 | 2.61e-74 | 67 | 426 | 108 | 465 |
| ANV85505.1 | 3.40e-72 | 67 | 426 | 106 | 463 |
| AFZ12267.1 | 4.20e-72 | 23 | 426 | 61 | 455 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5EJ1_A | 9.95e-13 | 67 | 299 | 128 | 380 | ChainA, Putative cellulose synthase [Cereibacter sphaeroides 2.4.1] |
| 4HG6_A | 1.05e-12 | 67 | 299 | 140 | 392 | ChainA, Cellulose Synthase Subunit A [Cereibacter sphaeroides] |
| 4P00_A | 1.05e-12 | 67 | 299 | 141 | 393 | ChainA, Cellulose Synthase A subunit [Cereibacter sphaeroides 2.4.1],4P02_A Chain A, Cellulose Synthase subunit A [Cereibacter sphaeroides 2.4.1],5EIY_A Chain A, Putative cellulose synthase [Cereibacter sphaeroides 2.4.1],5EJZ_A Chain A, Putative cellulose synthase [Cereibacter sphaeroides 2.4.1] |
| 7LBY_A | 5.90e-12 | 55 | 296 | 261 | 503 | ChainA, Cellulose synthase catalytic subunit [UDP-forming] [Escherichia coli K-12] |
| 6P61_A | 1.26e-08 | 67 | 171 | 13 | 113 | Structureof a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_B Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_C Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_D Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q3MB01 | 1.19e-67 | 67 | 426 | 106 | 463 | Beta-monoglucosyldiacylglycerol synthase OS=Trichormus variabilis (strain ATCC 29413 / PCC 7937) OX=240292 GN=Ava_2217 PE=1 SV=1 |
| Q8YMK0 | 1.19e-67 | 67 | 426 | 106 | 463 | Beta-monoglucosyldiacylglycerol synthase OS=Nostoc sp. (strain PCC 7120 / SAG 25.82 / UTEX 2576) OX=103690 GN=all4933 PE=1 SV=1 |
| P74165 | 8.12e-62 | 14 | 427 | 60 | 467 | Beta-monoglucosyldiacylglycerol synthase OS=Synechocystis sp. (strain PCC 6803 / Kazusa) OX=1111708 GN=sll1377 PE=1 SV=1 |
| P96587 | 4.30e-30 | 32 | 424 | 12 | 411 | Uncharacterized glycosyltransferase YdaM OS=Bacillus subtilis (strain 168) OX=224308 GN=ydaM PE=3 SV=1 |
| Q5HKQ0 | 3.13e-28 | 36 | 315 | 14 | 292 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=icaA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000027 | 0.000002 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
TMHMM Annotations download full data without filtering help
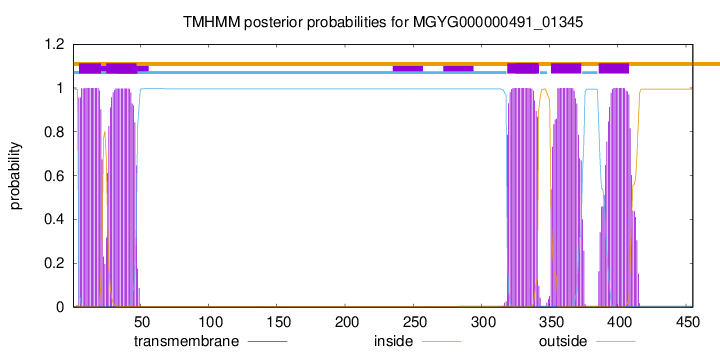
| start | end |
|---|---|
| 5 | 21 |
| 25 | 47 |
| 319 | 341 |
| 351 | 373 |
| 386 | 408 |
