You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000488_00658
You are here: Home > Sequence: MGYG000000488_00658
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Anaeroplasma sp900767915 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Acholeplasmatales; Anaeroplasmataceae; Anaeroplasma; Anaeroplasma sp900767915 | |||||||||||
| CAZyme ID | MGYG000000488_00658 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 52420; End: 56451 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 800 | 993 | 1.7e-31 | 0.8981481481481481 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01915 | Glyco_hydro_3_C | 4.12e-14 | 65 | 340 | 2 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| PLN03080 | PLN03080 | 4.33e-13 | 38 | 542 | 377 | 773 | Probable beta-xylosidase; Provisional |
| pfam14310 | Fn3-like | 2.36e-11 | 461 | 541 | 2 | 71 | Fibronectin type III-like domain. This domain has a fibronectin type III-like structure. It is often found in association with pfam00933 and pfam01915. Its function is unknown. |
| PRK15098 | PRK15098 | 2.68e-07 | 394 | 512 | 640 | 734 | beta-glucosidase BglX. |
| pfam05345 | He_PIG | 1.60e-04 | 1072 | 1129 | 11 | 76 | Putative Ig domain. This alignment represents the conserved core region of ~90 residue repeat found in several haemagglutinins and other cell surface proteins. Sequence similarities to (pfam02494) and (pfam00801) suggest an Ig-like fold (personal obs:C. Yeats). So this family may be similar in function to the (pfam02639) and (pfam02638) domains. This domain is also found in the WisP family of proteins of Tropheryma whipplei. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| VEU80230.1 | 1.39e-149 | 37 | 1257 | 28 | 1123 |
| QOS39237.1 | 7.91e-145 | 37 | 1225 | 46 | 1130 |
| VEU80232.1 | 9.57e-135 | 38 | 1063 | 66 | 919 |
| QRA08612.1 | 6.96e-110 | 47 | 1055 | 118 | 924 |
| QWS31125.1 | 6.96e-110 | 47 | 1055 | 118 | 924 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5WUG_A | 3.06e-35 | 49 | 971 | 33 | 747 | Expression,characterization and crystal structure of a novel beta-glucosidase from Paenibacillus barengoltzii [Paenibacillus barengoltzii],5WVP_A Expression, characterization and crystal structure of a novel beta-glucosidase from Paenibacillus barengoltzii [Paenibacillus barengoltzii] |
| 2X40_A | 6.60e-20 | 807 | 975 | 78 | 246 | Structureof beta-glucosidase 3B from Thermotoga neapolitana in complex with glycerol [Thermotoga neapolitana DSM 4359],2X41_A Structure of beta-glucosidase 3B from Thermotoga neapolitana in complex with glucose [Thermotoga neapolitana DSM 4359] |
| 2X42_A | 5.96e-19 | 807 | 975 | 78 | 246 | Structureof beta-glucosidase 3B from Thermotoga neapolitana in complex with alpha-D-glucose [Thermotoga neapolitana DSM 4359] |
| 7MS2_A | 1.99e-16 | 763 | 997 | 7 | 253 | ChainA, Thermostable beta-glucosidase B [Acetivibrio thermocellus AD2],7MS2_B Chain B, Thermostable beta-glucosidase B [Acetivibrio thermocellus AD2] |
| 6JXG_A | 1.93e-09 | 59 | 531 | 329 | 698 | CrystaslStructure of Beta-glucosidase D2-BGL from Chaetomella Raphigera [Chaetomella raphigera] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P16084 | 1.12e-46 | 53 | 971 | 29 | 769 | Beta-glucosidase A OS=Butyrivibrio fibrisolvens OX=831 GN=bglA PE=3 SV=1 |
| P15885 | 4.11e-38 | 53 | 971 | 10 | 696 | Beta-glucosidase OS=Ruminococcus albus OX=1264 PE=3 SV=1 |
| P27034 | 1.26e-18 | 762 | 973 | 3 | 224 | Beta-glucosidase OS=Rhizobium radiobacter OX=358 GN=cbg-1 PE=3 SV=1 |
| Q5B6C7 | 1.97e-17 | 798 | 984 | 45 | 237 | Probable beta-glucosidase H OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglH PE=3 SV=2 |
| A1DNN8 | 2.04e-17 | 754 | 994 | 7 | 252 | Probable beta-glucosidase J OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=bglJ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
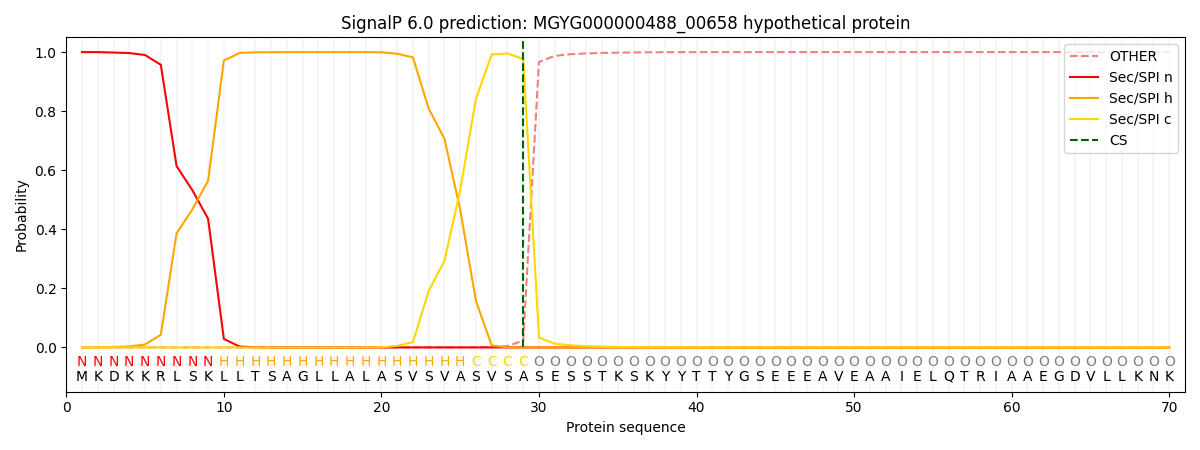
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000269 | 0.999068 | 0.000175 | 0.000170 | 0.000157 | 0.000140 |

