You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000387_01515
You are here: Home > Sequence: MGYG000000387_01515
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; ; | |||||||||||
| CAZyme ID | MGYG000000387_01515 | |||||||||||
| CAZy Family | GH95 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8711; End: 11977 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH95 | 109 | 921 | 5.6e-224 | 0.9653739612188366 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam14498 | Glyco_hyd_65N_2 | 1.19e-49 | 115 | 390 | 13 | 231 | Glycosyl hydrolase family 65, N-terminal domain. This domain represents a domain found to the N-terminus of the glycosyl hydrolase 65 family catalytic domain. |
| cd14256 | Dockerin_I | 2.04e-13 | 1025 | 1081 | 1 | 57 | Type I dockerin repeat domain. Bacterial cohesin domains bind to a complementary protein domain named dockerin, and this interaction is required for the formation of the cellulosome, a cellulose-degrading complex. The cellulosome consists of scaffoldin, a noncatalytic scaffolding polypeptide, that comprises repeating cohesion modules and a single carbohydrate-binding module (CBM). Specific calcium-dependent interactions between cohesins and dockerins appear to be essential for cellulosome assembly. This subfamily represents type I dockerins, which are responsible for anchoring a variety of enzymatic domains to the complex. |
| cd14253 | Dockerin | 2.76e-10 | 1026 | 1081 | 1 | 56 | Dockerin repeat domain. Dockerins are modules in the cellulosome complex that often anchor catalytic subunits by binding to cohesin domains of scaffolding proteins. Three types of dockerins and their corresponding cohesin have been described in the literature. This alignment models two consecutive dockerin repeats, the functional unit. |
| pfam00404 | Dockerin_1 | 6.89e-10 | 1026 | 1079 | 1 | 54 | Dockerin type I repeat. The dockerin repeat is the binding partner of the cohesin domain pfam00963. The cohesin-dockerin interaction is the crucial interaction for complex formation in the cellulosome. The dockerin repeats, each bearing homology to the EF-hand calcium-binding loop bind calcium. |
| cd14252 | Dockerin_like | 2.18e-06 | 1026 | 1081 | 1 | 57 | Dockerin repeat domains and domains resembling dockerin repeats. Dockerins are modules in the cellulosome complex that often anchor catalytic subunits by binding to cohesin domains of scaffolding proteins. Three types of dockerins and their corresponding cohesin have been described in the literature. This alignment models two consecutive dockerin repeats, the functional unit. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QPA42690.1 | 1.25e-170 | 111 | 944 | 58 | 789 |
| QPA36938.1 | 1.25e-170 | 111 | 944 | 58 | 789 |
| QHH85405.1 | 1.57e-170 | 111 | 944 | 58 | 789 |
| QRH05335.1 | 1.57e-170 | 111 | 944 | 58 | 789 |
| QHG35690.1 | 1.57e-170 | 111 | 944 | 58 | 789 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2EAB_A | 1.04e-138 | 107 | 916 | 43 | 844 | Crystalstructure of 1,2-a-L-fucosidase from Bifidobacterium bifidum (apo form) [Bifidobacterium bifidum],2EAB_B Crystal structure of 1,2-a-L-fucosidase from Bifidobacterium bifidum (apo form) [Bifidobacterium bifidum],2EAC_A Crystal structure of 1,2-a-L-fucosidase from Bifidobacterium bifidum in complex with deoxyfuconojirimycin [Bifidobacterium bifidum],2EAC_B Crystal structure of 1,2-a-L-fucosidase from Bifidobacterium bifidum in complex with deoxyfuconojirimycin [Bifidobacterium bifidum] |
| 2EAD_A | 7.72e-138 | 107 | 916 | 43 | 844 | ChainA, Alpha-fucosidase [Bifidobacterium bifidum],2EAD_B Chain B, Alpha-fucosidase [Bifidobacterium bifidum] |
| 2EAE_A | 1.47e-137 | 107 | 916 | 42 | 843 | ChainA, Alpha-fucosidase [Bifidobacterium bifidum] |
| 2RDY_A | 6.40e-102 | 110 | 958 | 12 | 790 | ChainA, BH0842 protein [Halalkalibacterium halodurans C-125],2RDY_B Chain B, BH0842 protein [Halalkalibacterium halodurans C-125] |
| 4UFC_A | 3.71e-99 | 110 | 919 | 32 | 739 | Crystalstructure of the GH95 enzyme BACOVA_03438 [Bacteroides ovatus],4UFC_B Crystal structure of the GH95 enzyme BACOVA_03438 [Bacteroides ovatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8L7W8 | 8.14e-80 | 115 | 903 | 66 | 794 | Alpha-L-fucosidase 2 OS=Arabidopsis thaliana OX=3702 GN=FUC95A PE=1 SV=1 |
| A2R797 | 1.82e-72 | 102 | 903 | 25 | 758 | Probable alpha-fucosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=afcA PE=3 SV=1 |
| Q5AU81 | 6.20e-60 | 100 | 918 | 24 | 793 | Alpha-fucosidase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=afcA PE=1 SV=1 |
| Q2USL3 | 1.06e-47 | 110 | 921 | 25 | 715 | Probable alpha-fucosidase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=afcA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
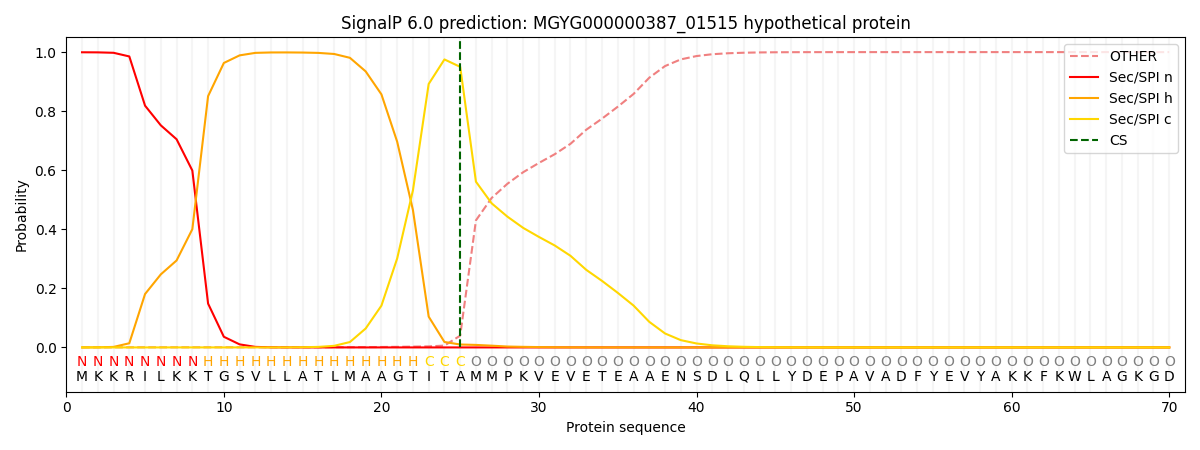
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001154 | 0.997710 | 0.000474 | 0.000263 | 0.000189 | 0.000175 |
