You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000265_03852
You are here: Home > Sequence: MGYG000000265_03852
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides nordii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides nordii | |||||||||||
| CAZyme ID | MGYG000000265_03852 | |||||||||||
| CAZy Family | PL6 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 40048; End: 41484 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL6 | 39 | 387 | 1.6e-107 | 0.9567723342939481 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd14251 | PL-6 | 1.62e-154 | 38 | 410 | 12 | 367 | Polysaccharide Lyase Family 6. Polysaccharide Lyase Family 6 is a family of beta-helical polysaccharide lyases. Members include alginate lyase (EC 4.2.2.3) and chondroitinase B (EC 4.2.2.19). Chondroitinase B is an enzyme that only cleaves the beta-(1,4)-linkage of dermatan sulfate (DS), leading to 4,5-unsaturated dermatan sulfate disaccharides as the product. DS is a highly sulfated, unbranched polysaccharide belonging to a family of glycosaminoglycans (GAGs) composed of alternating hexosamine (gluco- or galactosamine) and uronic acid (D-glucuronic or L-iduronic acid) moieties. DS contains alternating 1,4-beta-D-galactosamine (GalNac) and 1,3-alpha-L-iduronic acid units. The related chondroitin sulfate (CS) contains alternating GalNac and 1,3-beta-D-glucuronic acid units. Alginate lyases (known as either mannuronate (EC 4.2.2.3) or guluronate lyases (EC 4.2.2.11) catalyze the degradation of alginate, a copolymer of alpha-L-guluronate and its C5 epimer beta-D-mannuronate. |
| pfam14592 | Chondroitinas_B | 5.27e-89 | 28 | 446 | 4 | 423 | Chondroitinase B. This family includes chondroitinases. These enzymes cleave the glycosaminoglycan dermatan sulfate. |
| pfam13229 | Beta_helix | 7.55e-05 | 231 | 321 | 32 | 120 | Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDO71143.1 | 6.70e-208 | 1 | 461 | 2 | 461 |
| ALJ58476.1 | 5.28e-207 | 4 | 461 | 7 | 460 |
| QUT90411.1 | 8.66e-206 | 4 | 461 | 7 | 460 |
| QDO71096.1 | 6.36e-162 | 9 | 422 | 13 | 423 |
| ARV16326.1 | 4.52e-83 | 43 | 427 | 42 | 407 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6QPS_A | 4.07e-206 | 19 | 461 | 22 | 463 | Structuralcharacterization of a mannuronic acid specific polysaccharide family 6 lyase enzyme from human gut microbiota [Bacteroides cellulosilyticus],6QPS_B Structural characterization of a mannuronic acid specific polysaccharide family 6 lyase enzyme from human gut microbiota [Bacteroides cellulosilyticus] |
| 5GKD_A | 1.41e-68 | 39 | 427 | 17 | 388 | Structureof PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis],5GKD_B Structure of PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis],5GKD_C Structure of PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis],5GKD_D Structure of PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis] |
| 7O84_A | 1.94e-68 | 39 | 386 | 31 | 357 | ChainA, Alginate lyase [Pseudopedobacter saltans DSM 12145],7O84_B Chain B, Alginate lyase [Pseudopedobacter saltans DSM 12145] |
| 7O7A_A | 1.99e-68 | 39 | 386 | 31 | 357 | ChainA, Aliginate lyase [Pseudopedobacter saltans DSM 12145],7O7A_B Chain B, Aliginate lyase [Pseudopedobacter saltans DSM 12145] |
| 5GKQ_A | 1.02e-67 | 39 | 427 | 17 | 388 | Structureof PL6 family alginate lyase AlyGC mutant-R241A [Paraglaciecola chathamensis S18K6],5GKQ_B Structure of PL6 family alginate lyase AlyGC mutant-R241A [Paraglaciecola chathamensis S18K6] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q06365 | 1.08e-61 | 41 | 387 | 1 | 326 | Alginate lyase OS=Pseudomonas sp. (strain OS-ALG-9) OX=86038 GN=aly PE=3 SV=1 |
| Q46079 | 1.87e-22 | 4 | 310 | 11 | 308 | Chondroitinase-B OS=Pedobacter heparinus (strain ATCC 13125 / DSM 2366 / CIP 104194 / JCM 7457 / NBRC 12017 / NCIMB 9290 / NRRL B-14731 / HIM 762-3) OX=485917 GN=cslB PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
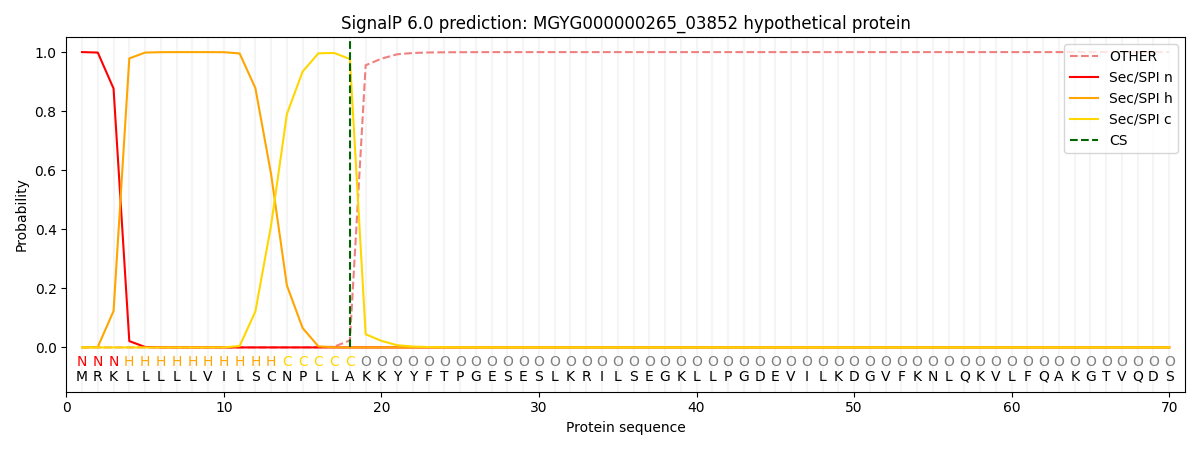
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000278 | 0.999117 | 0.000178 | 0.000138 | 0.000143 | 0.000133 |
