You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000238_02833
You are here: Home > Sequence: MGYG000000238_02833
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
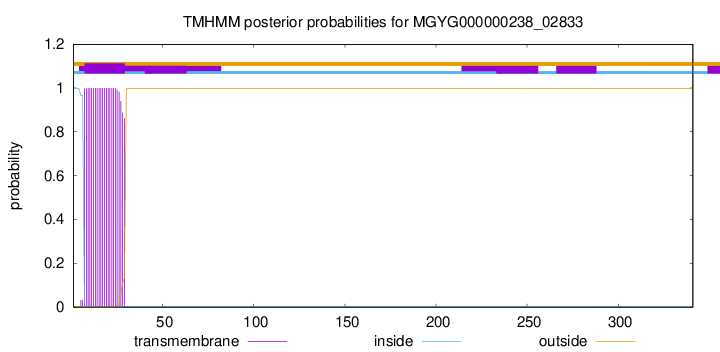
TMHMM annotations
Basic Information help
| Species | Enterococcus_D casseliflavus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Enterococcaceae; Enterococcus_D; Enterococcus_D casseliflavus | |||||||||||
| CAZyme ID | MGYG000000238_02833 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 41297; End: 42322 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16891 | CwlT-like | 3.67e-70 | 57 | 208 | 1 | 151 | CwlT-like N-terminal lysozyme domain and similar domains. CwlT is a bifunctional cell wall hydrolase containing an N-terminal lysozyme domain and a C-terminal NlpC/P60 endopeptidase domain (gamma-d-D-glutamyl-L-diamino acid endopeptidase), and has been implicated in the spread of transposons. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| pfam13702 | Lysozyme_like | 8.89e-68 | 51 | 207 | 1 | 165 | Lysozyme-like. |
| pfam00877 | NLPC_P60 | 3.70e-38 | 232 | 338 | 1 | 105 | NlpC/P60 family. The function of this domain is unknown. It is found in several lipoproteins. |
| PRK13914 | PRK13914 | 3.77e-35 | 212 | 339 | 358 | 481 | invasion associated endopeptidase. |
| COG0791 | Spr | 1.24e-29 | 166 | 336 | 25 | 196 | Cell wall-associated hydrolase, NlpC family [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QKG73966.1 | 8.55e-157 | 1 | 341 | 1 | 339 |
| QZO09928.1 | 8.55e-157 | 1 | 341 | 1 | 339 |
| QOG31972.1 | 8.55e-157 | 1 | 341 | 1 | 339 |
| QOG26341.1 | 8.55e-157 | 1 | 341 | 1 | 339 |
| AEP33213.1 | 8.55e-157 | 1 | 341 | 1 | 339 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4HPE_A | 1.47e-127 | 50 | 341 | 20 | 308 | ChainA, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_B Chain B, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_C Chain C, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_D Chain D, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_E Chain E, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_F Chain F, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630] |
| 4FDY_A | 2.91e-121 | 50 | 339 | 22 | 310 | ChainA, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50],4FDY_B Chain B, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50] |
| 7CFL_A | 1.39e-16 | 232 | 339 | 26 | 137 | ChainA, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_B Chain B, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_C Chain C, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_D Chain D, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile] |
| 2HBW_A | 1.50e-13 | 211 | 341 | 85 | 228 | ChainA, NLP/P60 protein [Trichormus variabilis ATCC 29413] |
| 2K1G_A | 4.73e-12 | 222 | 339 | 8 | 123 | SolutionNMR structure of lipoprotein spr from Escherichia coli K12. Northeast Structural Genomics target ER541-37-162 [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P96645 | 4.99e-79 | 56 | 340 | 59 | 329 | Probable endopeptidase YddH OS=Bacillus subtilis (strain 168) OX=224308 GN=yddH PE=3 SV=1 |
| P0DC91 | 9.98e-28 | 53 | 211 | 29 | 194 | Pneumococcal vaccine antigen A homolog OS=Streptococcus pyogenes serotype M3 (strain SSI-1) OX=193567 GN=pvaA PE=3 SV=1 |
| P0DC90 | 9.98e-28 | 53 | 211 | 29 | 194 | Pneumococcal vaccine antigen A homolog OS=Streptococcus pyogenes serotype M3 (strain ATCC BAA-595 / MGAS315) OX=198466 GN=pvaA PE=3 SV=1 |
| Q5XC63 | 1.94e-27 | 53 | 211 | 29 | 194 | Pneumococcal vaccine antigen A homolog OS=Streptococcus pyogenes serotype M6 (strain ATCC BAA-946 / MGAS10394) OX=286636 GN=pvaA PE=3 SV=1 |
| Q8P120 | 1.94e-27 | 53 | 211 | 29 | 194 | Pneumococcal vaccine antigen A homolog OS=Streptococcus pyogenes serotype M18 (strain MGAS8232) OX=186103 GN=pvaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
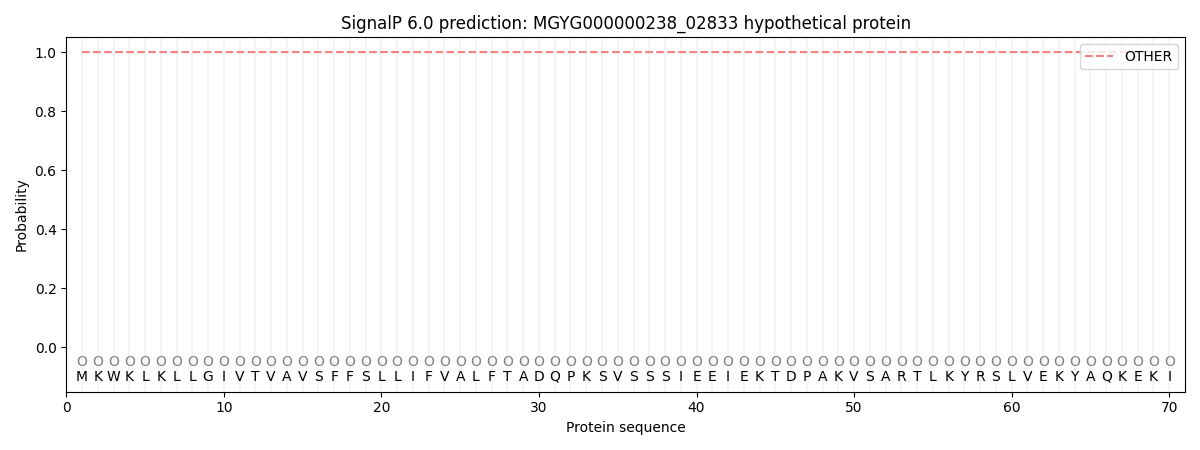
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000012 | 0.000012 | 0.000001 | 0.000000 | 0.000000 | 0.000001 |