You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000174_00250
You are here: Home > Sequence: MGYG000000174_00250
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides faecis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides faecis | |||||||||||
| CAZyme ID | MGYG000000174_00250 | |||||||||||
| CAZy Family | GH33 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 302436; End: 304067 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 167 | 531 | 1.4e-88 | 0.9385964912280702 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 4.62e-87 | 171 | 530 | 8 | 328 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| pfam13859 | BNR_3 | 9.33e-27 | 179 | 435 | 1 | 213 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
| pfam14873 | BNR_assoc_N | 9.18e-24 | 21 | 148 | 1 | 148 | N-terminal domain of BNR-repeat neuraminidase. This domain is usually found at the N-terminus of the BNR-repeat neuraminidase protein family. |
| pfam13088 | BNR_2 | 3.85e-15 | 188 | 522 | 1 | 280 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
| COG4409 | NanH | 1.38e-10 | 173 | 525 | 271 | 643 | Neuraminidase (sialidase) [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| SCD21035.1 | 5.92e-275 | 1 | 541 | 1 | 539 |
| QEW37925.1 | 6.57e-272 | 26 | 542 | 50 | 567 |
| BBD46000.1 | 7.73e-264 | 8 | 541 | 8 | 539 |
| CBW24233.1 | 5.69e-256 | 26 | 542 | 3 | 520 |
| QRP88498.1 | 9.23e-256 | 22 | 542 | 22 | 543 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6MYV_A | 2.50e-109 | 21 | 543 | 1 | 522 | Sialidase26co-crystallized with DANA-Gc [bacterium],6MYV_B Sialidase26 co-crystallized with DANA-Gc [bacterium],6MYV_C Sialidase26 co-crystallized with DANA-Gc [bacterium],6MYV_D Sialidase26 co-crystallized with DANA-Gc [bacterium] |
| 6MRV_A | 5.47e-109 | 21 | 543 | 22 | 543 | Sialidase26co-crystallized with DANA [bacterium],6MRV_B Sialidase26 co-crystallized with DANA [bacterium],6MRX_A Sialidase26 apo [bacterium],6MRX_B Sialidase26 apo [bacterium],6MRX_C Sialidase26 apo [bacterium],6MRX_D Sialidase26 apo [bacterium] |
| 4Q6K_A | 2.48e-102 | 21 | 543 | 4 | 525 | Crystalstructure of a putative neuraminidase (BACCAC_01090) from Bacteroides caccae ATCC 43185 at 1.90 A resolution (PSI Community Target) [Bacteroides caccae ATCC 43185],4Q6K_B Crystal structure of a putative neuraminidase (BACCAC_01090) from Bacteroides caccae ATCC 43185 at 1.90 A resolution (PSI Community Target) [Bacteroides caccae ATCC 43185],4Q6K_C Crystal structure of a putative neuraminidase (BACCAC_01090) from Bacteroides caccae ATCC 43185 at 1.90 A resolution (PSI Community Target) [Bacteroides caccae ATCC 43185],4Q6K_D Crystal structure of a putative neuraminidase (BACCAC_01090) from Bacteroides caccae ATCC 43185 at 1.90 A resolution (PSI Community Target) [Bacteroides caccae ATCC 43185] |
| 4FJ6_A | 9.25e-102 | 22 | 542 | 3 | 521 | Crystalstructure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503],4FJ6_B Crystal structure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503],4FJ6_C Crystal structure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503],4FJ6_D Crystal structure of a glycoside hydrolase family 33, candidate sialidase (BDI_2946) from Parabacteroides distasonis ATCC 8503 at 1.90 A resolution [Parabacteroides distasonis ATCC 8503] |
| 4BBW_A | 3.74e-96 | 21 | 542 | 23 | 543 | Thecrystal structure of Sialidase VPI 5482 (BTSA) from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P31206 | 1.48e-109 | 22 | 543 | 24 | 544 | Sialidase OS=Bacteroides fragilis (strain YCH46) OX=295405 GN=nanH PE=3 SV=2 |
| Q02834 | 3.49e-29 | 171 | 525 | 61 | 386 | Sialidase OS=Micromonospora viridifaciens OX=1881 GN=nedA PE=1 SV=1 |
| P15698 | 1.43e-24 | 178 | 527 | 55 | 382 | Sialidase OS=Paeniclostridium sordellii OX=1505 PE=1 SV=1 |
| P29767 | 8.22e-24 | 173 | 525 | 391 | 809 | Sialidase OS=Clostridium septicum OX=1504 PE=3 SV=1 |
| P10481 | 2.91e-20 | 178 | 527 | 37 | 364 | Sialidase OS=Clostridium perfringens OX=1502 GN=nanH PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
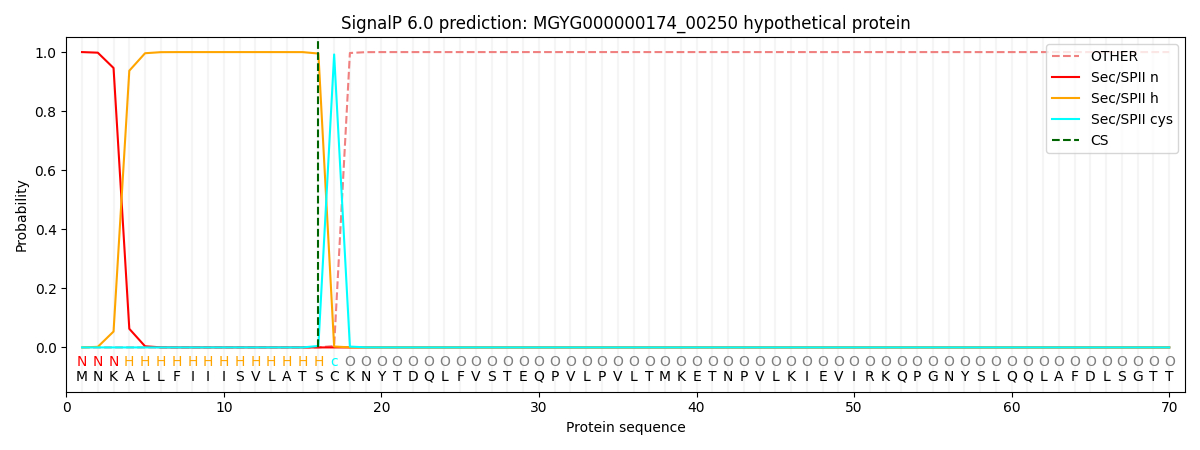
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000001 | 1.000060 | 0.000000 | 0.000000 | 0.000000 |
