You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000123_04832
You are here: Home > Sequence: MGYG000000123_04832
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Blautia sp001304935 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Blautia; Blautia sp001304935 | |||||||||||
| CAZyme ID | MGYG000000123_04832 | |||||||||||
| CAZy Family | GH112 | |||||||||||
| CAZyme Description | 1,3-beta-galactosyl-N-acetylhexosamine phosphorylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4886; End: 7054 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH112 | 9 | 721 | 0 | 0.9986013986013986 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| TIGR02336 | TIGR02336 | 0.0 | 6 | 721 | 1 | 719 | 1,3-beta-galactosyl-N-acetylhexosamine phosphorylase. Members of this family are found in phylogenetically diverse bacteria, including Clostridium perfringens (in the Firmicutes), Bifidobacterium longum and Propionibacterium acnes (in the Actinobacteria), and Vibrio vulnificus (in the Proteobacteria), most of which occur as mammalian pathogens or commensals. The nominal activity, 1,3-beta-galactosyl-N-acetylhexosamine phosphorylase (EC 2.4.1.211), varies somewhat from instance to instance in relative rates for closely related substrates. [Energy metabolism, Biosynthesis and degradation of polysaccharides] |
| pfam09508 | Lact_bio_phlase | 0.0 | 9 | 442 | 1 | 434 | Lacto-N-biose phosphorylase N-terminal TIM barrel domain. The gene which codes for this protein in gut-bacteria is located in a novel putative operon for galactose metabolism. The protein appears to be a carbohydrate-processing phosphorolytic enzyme (EC:2.4.1.211), unlike either glycoside hydrolases or glycoside lyase. Intestinal colonisation by bifidobacteria is important for human health, especially in pediatrics, because colonisation seems to prevent infection by some pathogenic bacteria that cause diarrhoea or other illnesses. The operon seems to be involved in intestinal colonisation by bifidobacteria mediated by metabolism of mucin sugars. In addition, it may also resolve the question of the nature of the bifidus factor in human milk as the lacto-N-biose structure found in milk oligosaccharides. |
| pfam17385 | LBP_M | 1.41e-135 | 444 | 663 | 1 | 221 | Lacto-N-biose phosphorylase central domain. The gene which codes for this protein in gut-bacteria is located in a novel putative operon for galactose metabolism. The protein appears to be a carbohydrate-processing phosphorolytic enzyme (EC:2.4.1.211), unlike either glycoside hydrolases or glycoside lyase. Intestinal colonisation by bifidobacteria is important for human health, especially in pediatrics, because colonisation seems to prevent infection by some pathogenic bacteria that cause diarrhoea or other illnesses. The operon seems to be involved in intestinal colonisation by bifidobacteria mediated by metabolism of mucin sugars. In addition, it may also resolve the question of the nature of the bifidus factor in human milk as the lacto-N-biose structure found in milk oligosaccharides. |
| pfam17386 | LBP_C | 1.68e-23 | 668 | 720 | 1 | 53 | Lacto-N-biose phosphorylase C-terminal domain. The gene which codes for this protein in gut-bacteria is located in a novel putative operon for galactose metabolism. The protein appears to be a carbohydrate-processing phosphorolytic enzyme (EC:2.4.1.211), unlike either glycoside hydrolases or glycoside lyase. Intestinal colonisation by bifidobacteria is important for human health, especially in pediatrics, because colonisation seems to prevent infection by some pathogenic bacteria that cause diarrhoea or other illnesses. The operon seems to be involved in intestinal colonisation by bifidobacteria mediated by metabolism of mucin sugars. In addition, it may also resolve the question of the nature of the bifidus factor in human milk as the lacto-N-biose structure found in milk oligosaccharides. |
| cd10950 | CE4_BsYlxY_like | 3.48e-06 | 47 | 177 | 28 | 134 | Putative catalytic NodB homology domain of uncharacterized protein YlxY from Bacillus subtilis and its bacterial homologs. The Bacillus subtilis genome contains six polysaccharide deacetylase gene homologs: pdaA, pdaB (previously known as ybaN), yheN, yjeA, yxkH and ylxY. This family is represented by Bacillus subtilis putative polysaccharide deacetylase BsYlxY, encoded by the ylxY gene, which is a member of the carbohydrate esterase 4 (CE4) superfamily. Although its biological function still remains unknown, BsYlxY shows high sequence homology to the catalytic domain of Bacillus subtilis pdaB gene encoding a putative polysaccharide deacetylase (BsPdaB), which is essential for the maintenance of spores after the late stage of sporulation and is highly conserved in spore-forming bacteria. However, disruption of the ylxY gene in B. subtilis did not cause any sporulation defect. Moreover, the Asp residue in the classical His-His-Asp zinc-binding motif of CE4 esterases is mutated to a Val residue in this family. Other catalytically relevant residues of CE4 esterases are also not conserved, which suggest that members of this family may be inactive. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QBE97492.1 | 0.0 | 1 | 722 | 1 | 722 |
| QIB58571.1 | 0.0 | 7 | 722 | 1 | 716 |
| QMW81461.1 | 0.0 | 7 | 722 | 1 | 716 |
| QQQ93893.1 | 0.0 | 1 | 722 | 1 | 722 |
| ANU76318.1 | 0.0 | 1 | 722 | 1 | 722 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3WFZ_A | 1.77e-233 | 9 | 721 | 5 | 750 | Crystalstructure of Galacto-N-Biose/Lacto-N-Biose I Phosphorylase C236Y Mutant [Bifidobacterium longum subsp. longum JCM 1217],3WFZ_B Crystal structure of Galacto-N-Biose/Lacto-N-Biose I Phosphorylase C236Y Mutant [Bifidobacterium longum subsp. longum JCM 1217],3WFZ_C Crystal structure of Galacto-N-Biose/Lacto-N-Biose I Phosphorylase C236Y Mutant [Bifidobacterium longum subsp. longum JCM 1217],3WFZ_D Crystal structure of Galacto-N-Biose/Lacto-N-Biose I Phosphorylase C236Y Mutant [Bifidobacterium longum subsp. longum JCM 1217] |
| 2ZUS_A | 4.05e-232 | 9 | 721 | 5 | 750 | Crystalstructure of Galacto-N-biose/Lacto-N-biose I phosphorylase [Bifidobacterium longum],2ZUS_B Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase [Bifidobacterium longum],2ZUS_C Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase [Bifidobacterium longum],2ZUS_D Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase [Bifidobacterium longum],2ZUT_A Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GalNAc [Bifidobacterium longum],2ZUT_B Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GalNAc [Bifidobacterium longum],2ZUT_C Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GalNAc [Bifidobacterium longum],2ZUT_D Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GalNAc [Bifidobacterium longum],2ZUU_A Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc [Bifidobacterium longum],2ZUU_B Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc [Bifidobacterium longum],2ZUU_C Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc [Bifidobacterium longum],2ZUU_D Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc [Bifidobacterium longum],2ZUV_A Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc, Ethylene glycol, and nitrate [Bifidobacterium longum],2ZUV_B Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc, Ethylene glycol, and nitrate [Bifidobacterium longum],2ZUW_A Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc and sulfate [Bifidobacterium longum],2ZUW_B Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc and sulfate [Bifidobacterium longum],2ZUW_C Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc and sulfate [Bifidobacterium longum],2ZUW_D Crystal structure of Galacto-N-biose/Lacto-N-biose I phosphorylase in complex with GlcNAc and sulfate [Bifidobacterium longum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A9KIW5 | 0.0 | 1 | 722 | 1 | 723 | 1,3-beta-galactosyl-N-acetylhexosamine phosphorylase Cphy0577 OS=Lachnoclostridium phytofermentans (strain ATCC 700394 / DSM 18823 / ISDg) OX=357809 GN=Cphy_0577 PE=1 SV=1 |
| A9KQ75 | 0.0 | 1 | 720 | 1 | 719 | 1,3-beta-galactosyl-N-acetylhexosamine phosphorylase Cphy3030 OS=Lachnoclostridium phytofermentans (strain ATCC 700394 / DSM 18823 / ISDg) OX=357809 GN=Cphy_3030 PE=1 SV=1 |
| E8MF13 | 1.70e-231 | 9 | 721 | 5 | 750 | 1,3-beta-galactosyl-N-acetylhexosamine phosphorylase OS=Bifidobacterium longum subsp. longum (strain ATCC 15707 / DSM 20219 / JCM 1217 / NCTC 11818 / E194b) OX=565042 GN=lnpA PE=1 SV=1 |
| A9KHK4 | 1.86e-137 | 3 | 700 | 4 | 700 | D-galactosyl-beta-1->4-L-rhamnose phosphorylase OS=Lachnoclostridium phytofermentans (strain ATCC 700394 / DSM 18823 / ISDg) OX=357809 GN=Cphy_1920 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
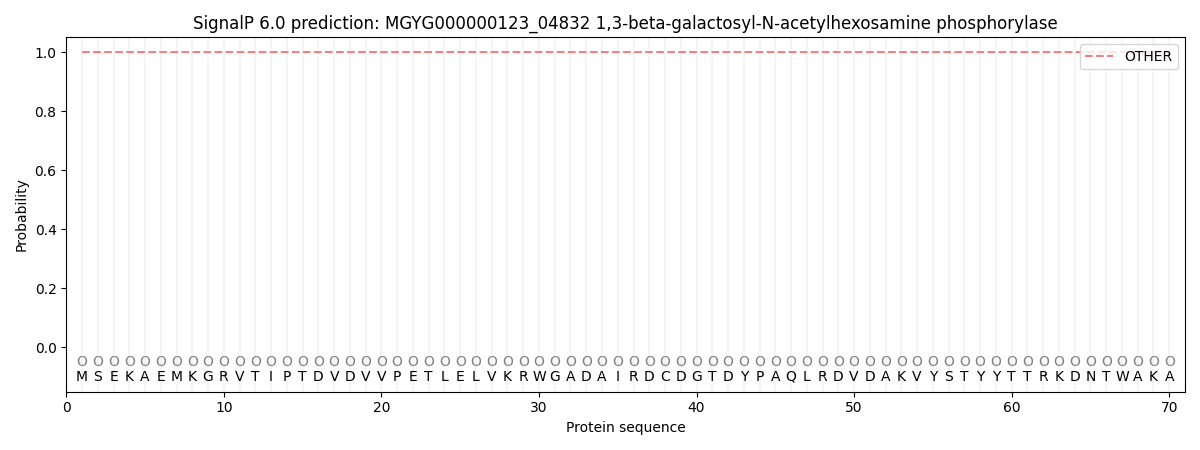
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000067 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
