You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000013_01838
You are here: Home > Sequence: MGYG000000013_01838
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp902362375 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp902362375 | |||||||||||
| CAZyme ID | MGYG000000013_01838 | |||||||||||
| CAZy Family | CE6 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 300083; End: 302341 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE6 | 581 | 672 | 2.4e-26 | 0.98989898989899 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03629 | SASA | 4.13e-75 | 521 | 743 | 2 | 223 | Carbohydrate esterase, sialic acid-specific acetylesterase. The catalytic triad of this esterase enzyme comprises residues Ser127, His403 and Asp391 in UniProtKB:P70665. |
| pfam03629 | SASA | 5.12e-14 | 103 | 398 | 2 | 223 | Carbohydrate esterase, sialic acid-specific acetylesterase. The catalytic triad of this esterase enzyme comprises residues Ser127, His403 and Asp391 in UniProtKB:P70665. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDH57420.1 | 0.0 | 1 | 750 | 1 | 750 |
| EDV06361.1 | 0.0 | 1 | 749 | 1 | 749 |
| QDO68497.1 | 0.0 | 1 | 749 | 1 | 749 |
| QRQ49244.1 | 0.0 | 1 | 752 | 1 | 752 |
| QUT44990.1 | 0.0 | 1 | 752 | 1 | 752 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7KMM_A | 3.77e-64 | 24 | 503 | 24 | 636 | ChainA, Sialic acid-specific 9-O-acetylesterase [Xanthomonas citri pv. citri str. 306],7KMM_B Chain B, Sialic acid-specific 9-O-acetylesterase [Xanthomonas citri pv. citri str. 306] |
| 2APJ_A | 4.14e-17 | 518 | 743 | 19 | 254 | X-RayStructure of Protein from Arabidopsis Thaliana AT4G34215 at 1.6 Angstrom Resolution [Arabidopsis thaliana],2APJ_B X-Ray Structure of Protein from Arabidopsis Thaliana AT4G34215 at 1.6 Angstrom Resolution [Arabidopsis thaliana],2APJ_C X-Ray Structure of Protein from Arabidopsis Thaliana AT4G34215 at 1.6 Angstrom Resolution [Arabidopsis thaliana],2APJ_D X-Ray Structure of Protein from Arabidopsis Thaliana AT4G34215 at 1.6 Angstrom Resolution [Arabidopsis thaliana] |
| 1ZMB_A | 6.78e-17 | 524 | 746 | 5 | 226 | CrystalStructure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_B Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_C Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_D Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_E Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_F Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5RFU0 | 1.68e-29 | 22 | 418 | 25 | 421 | Sialate O-acetylesterase OS=Pongo abelii OX=9601 GN=SIAE PE=2 SV=1 |
| P70665 | 2.00e-29 | 22 | 418 | 25 | 447 | Sialate O-acetylesterase OS=Mus musculus OX=10090 GN=Siae PE=1 SV=3 |
| Q9HAT2 | 2.26e-29 | 22 | 418 | 25 | 421 | Sialate O-acetylesterase OS=Homo sapiens OX=9606 GN=SIAE PE=1 SV=1 |
| P82450 | 3.62e-29 | 22 | 418 | 25 | 448 | Sialate O-acetylesterase OS=Rattus norvegicus OX=10116 GN=Siae PE=1 SV=2 |
| Q8L9J9 | 5.02e-17 | 518 | 743 | 19 | 254 | Probable carbohydrate esterase At4g34215 OS=Arabidopsis thaliana OX=3702 GN=At4g34215 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
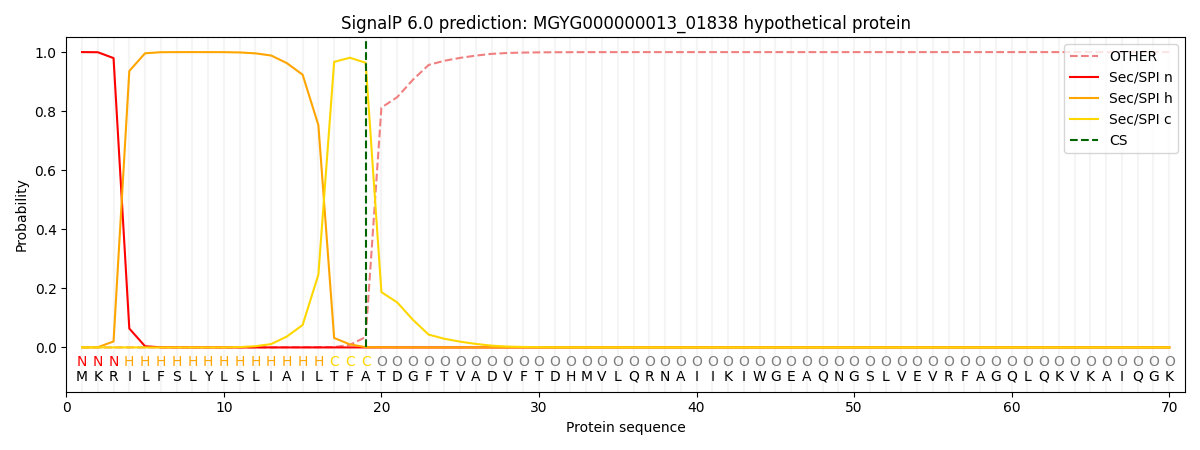
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000447 | 0.998860 | 0.000194 | 0.000161 | 0.000154 | 0.000160 |