You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000008_00175
You are here: Home > Sequence: MGYG000000008_00175
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Lactobacillus johnsonii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Lactobacillus; Lactobacillus johnsonii | |||||||||||
| CAZyme ID | MGYG000000008_00175 | |||||||||||
| CAZy Family | GT8 | |||||||||||
| CAZyme Description | Glycosyltransferase stabilizing protein Gtf2 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 170766; End: 172784 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT101 | 424 | 648 | 4.3e-93 | 0.9955357142857143 |
| GT8 | 3 | 229 | 5.8e-49 | 0.867704280155642 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| TIGR03728 | glyco_access_1 | 5.61e-157 | 407 | 672 | 1 | 265 | glycosyltransferase, SP_1767 family. Members of this protein family are putative glycosyltransferases. Some members are found close to genes for the accessory secretory (SecA2) system, and are suggested by Partial Phylogenetic Profiling to correlate with SecA2 systems. Glycosylation, therefore, may occur in the cytosol prior to secretion. |
| pfam08759 | GT-D | 6.87e-120 | 425 | 649 | 1 | 223 | Glycosyltransferase GT-D fold. This domain is found at the C-terminus of proteins such as the probable glycosyltransferase Gly that also contain the glycosyl transferase domain at the N-terminus. It is also found N-terminal in numerous putative glycosyltransferases such as GalT1. GalT1 has been shown to catalyze the third step of Fap1 glycosylation. This domain is structurally distinct from all known GT folds of glycosyltransferases and contains a metal binding site. This new glycosyltransferase fold has been named GT-D. |
| cd04194 | GT8_A4GalT_like | 1.53e-62 | 2 | 229 | 1 | 247 | A4GalT_like proteins catalyze the addition of galactose or glucose residues to the lipooligosaccharide (LOS) or lipopolysaccharide (LPS) of the bacterial cell surface. The members of this family of glycosyltransferases catalyze the addition of galactose or glucose residues to the lipooligosaccharide (LOS) or lipopolysaccharide (LPS) of the bacterial cell surface. The enzymes exhibit broad substrate specificities. The known functions found in this family include: Alpha-1,4-galactosyltransferase, LOS-alpha-1,3-D-galactosyltransferase, UDP-glucose:(galactosyl) LPS alpha1,2-glucosyltransferase, UDP-galactose: (glucosyl) LPS alpha1,2-galactosyltransferase, and UDP-glucose:(glucosyl) LPS alpha1,2-glucosyltransferase. Alpha-1,4-galactosyltransferase from N. meningitidis adds an alpha-galactose from UDP-Gal (the donor) to a terminal lactose (the acceptor) of the LOS structure of outer membrane. LOSs are virulence factors that enable the organism to evade the immune system of host cells. In E. coli, the three alpha-1,2-glycosyltransferases, that are involved in the synthesis of the outer core region of the LPS, are all members of this family. The three enzymes share 40 % of sequence identity, but have different sugar donor or acceptor specificities, representing the structural diversity of LPS. |
| COG1442 | RfaJ | 6.47e-47 | 2 | 310 | 3 | 315 | Lipopolysaccharide biosynthesis protein, LPS:glycosyltransferase [Cell wall/membrane/envelope biogenesis]. |
| pfam01501 | Glyco_transf_8 | 1.92e-32 | 4 | 227 | 2 | 247 | Glycosyl transferase family 8. This family includes enzymes that transfer sugar residues to donor molecules. Members of this family are involved in lipopolysaccharide biosynthesis and glycogen synthesis. This family includes Lipopolysaccharide galactosyltransferase, lipopolysaccharide glucosyltransferase 1, and glycogenin glucosyltransferase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AAS09481.1 | 0.0 | 6 | 672 | 2 | 668 |
| AOG26108.1 | 0.0 | 1 | 672 | 1 | 672 |
| AYN50091.1 | 0.0 | 1 | 672 | 1 | 672 |
| QXL47311.1 | 0.0 | 1 | 672 | 1 | 672 |
| QMT67762.1 | 0.0 | 1 | 672 | 1 | 672 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4PHR_A | 1.55e-92 | 405 | 671 | 12 | 276 | Domainof unknown function 1792 (DUF1792) with manganese [Streptococcus parasanguinis FW213] |
| 4PFX_A | 1.55e-92 | 405 | 671 | 12 | 276 | Thehighly conserved domain of unknown function 1792 has a distinct glycosyltransferase fold [Streptococcus parasanguinis FW213] |
| 5V4A_A | 3.91e-92 | 405 | 671 | 6 | 271 | ANew Glycosyltransferase (DUF1792) from Streptococcus sanguinis [Streptococcus sanguinis SK36],5V4A_B A New Glycosyltransferase (DUF1792) from Streptococcus sanguinis [Streptococcus sanguinis SK36] |
| 4PHS_A | 1.32e-90 | 405 | 671 | 12 | 276 | Selenomethioninesubstituted structure of domain of unknown function 1792 (DUF1792) [Streptococcus parasanguinis FW213] |
| 5GVW_A | 5.21e-80 | 2 | 378 | 6 | 386 | Crystalstructure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae],5GVW_B Crystal structure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae],5GVW_C Crystal structure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae],5GVW_D Crystal structure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9AEU2 | 1.29e-154 | 2 | 671 | 16 | 679 | Probable glycosyl transferase Gly OS=Streptococcus gordonii OX=1302 GN=gly PE=3 SV=2 |
| A0A0H2URB1 | 1.10e-153 | 2 | 671 | 3 | 812 | Glycosyltransferase GlyD OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=glyD PE=1 SV=1 |
| A0A0H2URJ6 | 2.85e-79 | 2 | 378 | 6 | 386 | Glycosyltransferase GlyE OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=glyE PE=1 SV=1 |
| A0A0H2URA3 | 1.71e-67 | 2 | 396 | 4 | 402 | Glycosyltransferase GlyB OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=glyB PE=3 SV=1 |
| A0A0H2URH2 | 3.00e-65 | 2 | 393 | 3 | 393 | Glycosyltransferase GlyF OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=glyF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
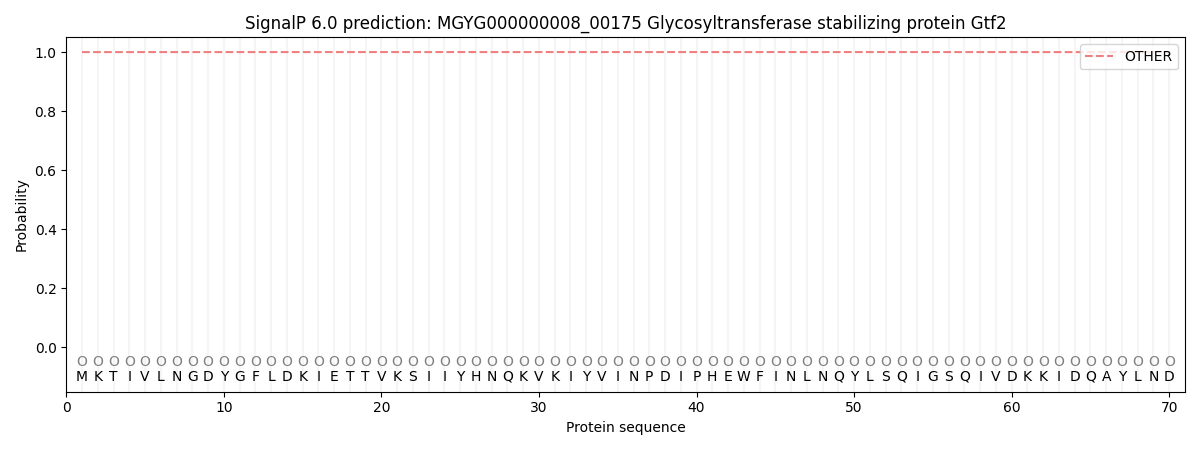
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000086 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
